| fit2d.C: example illustrating how to fit a 2-d histogram of type y=f(x) | Fitting tutorials | fitCircle.C: Generate points distributed with some errors around a circle |
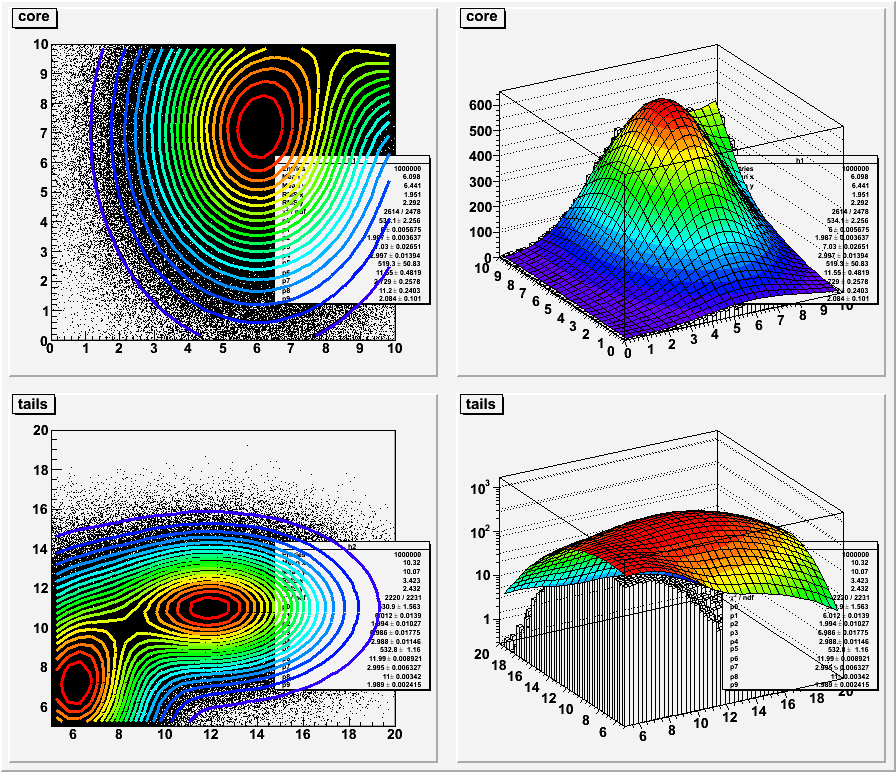
fit2dHist.C: Example to fit two histograms at the same time via TVirtualFitter
// //+ Example to fit two histograms at the same time via TVirtualFitter // // To execute this tutorial, you can do: // // root > .x fit2dHist.C (executing via CINT, slow) // or // root > .x fit2dHist.C+ (executing via ACLIC , fast, with Minuit) // root > .x fit2dHist.C+(2) (executing via ACLIC , fast, with Minuit2) // or using the option to fit independently the 2 histos // root > .x fit2dHist.C+(10) (via ACLIC, fast, independent fits with Minuit) // root > .x fit2dHist.C+(12) (via ACLIC, fast, independent fits with Minuit2) // // Note that you can also execute this script in batch with eg, // root -b -q "fit2dHist.C+(12)" // // or execute interactively from the shell // root fit2dHist.C+ // root "fit2dHist.C+(12)" // // Authors: Lorenzo Moneta, Rene Brun 18/01/2006 #include "TH2D.h" #include "TF2.h" #include "TCanvas.h" #include "TStyle.h" #include "TRandom3.h" #include "TVirtualFitter.h" #include "TList.h" #include <iostream> double gauss2D(double *x, double *par) { double z1 = double((x[0]-par[1])/par[2]); double z2 = double((x[1]-par[3])/par[4]); return par[0]*exp(-0.5*(z1*z1+z2*z2)); } double my2Dfunc(double *x, double *par) { return gauss2D(x,&par[0]) + gauss2D(x,&par[5]); } // data need to be globals to be visible by fcn TRandom3 rndm; TH2D *h1, *h2; Int_t npfits; void myFcn(Int_t & /*nPar*/, Double_t * /*grad*/ , Double_t &fval, Double_t *p, Int_t /*iflag */ ) { TAxis *xaxis1 = h1->GetXaxis(); TAxis *yaxis1 = h1->GetYaxis(); TAxis *xaxis2 = h2->GetXaxis(); TAxis *yaxis2 = h2->GetYaxis(); int nbinX1 = h1->GetNbinsX(); int nbinY1 = h1->GetNbinsY(); int nbinX2 = h2->GetNbinsX(); int nbinY2 = h2->GetNbinsY(); double chi2 = 0; double x[2]; double tmp; npfits = 0; for (int ix = 1; ix <= nbinX1; ++ix) { x[0] = xaxis1->GetBinCenter(ix); for (int iy = 1; iy <= nbinY1; ++iy) { if ( h1->GetBinError(ix,iy) > 0 ) { x[1] = yaxis1->GetBinCenter(iy); tmp = (h1->GetBinContent(ix,iy) - my2Dfunc(x,p))/h1->GetBinError(ix,iy); chi2 += tmp*tmp; npfits++; } } } for (int ix = 1; ix <= nbinX2; ++ix) { x[0] = xaxis2->GetBinCenter(ix); for (int iy = 1; iy <= nbinY2; ++iy) { if ( h2->GetBinError(ix,iy) > 0 ) { x[1] = yaxis2->GetBinCenter(iy); tmp = (h2->GetBinContent(ix,iy) - my2Dfunc(x,p))/h2->GetBinError(ix,iy); chi2 += tmp*tmp; npfits++; } } } fval = chi2; } void FillHisto(TH2D * h, int n, double * p) { const double mx1 = p[1]; const double my1 = p[3]; const double sx1 = p[2]; const double sy1 = p[4]; const double mx2 = p[6]; const double my2 = p[8]; const double sx2 = p[7]; const double sy2 = p[9]; //const double w1 = p[0]*sx1*sy1/(p[5]*sx2*sy2); const double w1 = 0.5; double x, y; for (int i = 0; i < n; ++i) { // generate randoms with larger gaussians rndm.Rannor(x,y); double r = rndm.Rndm(1); if (r < w1) { x = x*sx1 + mx1; y = y*sy1 + my1; } else { x = x*sx2 + mx2; y = y*sy2 + my2; } h->Fill(x,y); } } int fit2dHist(int option=1) { // create two histograms int nbx1 = 50; int nby1 = 50; int nbx2 = 50; int nby2 = 50; double xlow1 = 0.; double ylow1 = 0.; double xup1 = 10.; double yup1 = 10.; double xlow2 = 5.; double ylow2 = 5.; double xup2 = 20.; double yup2 = 20.; h1 = new TH2D("h1","core",nbx1,xlow1,xup1,nby1,ylow1,yup1); h2 = new TH2D("h2","tails",nbx2,xlow2,xup2,nby2,ylow2,yup2); double iniParams[10] = { 100, 6., 2., 7., 3, 100, 12., 3., 11., 2. }; // create fit function TF2 * func = new TF2("func",my2Dfunc,xlow2,xup2,ylow2,yup2, 10); func->SetParameters(iniParams); // fill Histos int n1 = 1000000; int n2 = 1000000; FillHisto(h1,n1,iniParams); FillHisto(h2,n2,iniParams); // scale histograms to same heights (for fitting) double dx1 = (xup1-xlow1)/double(nbx1); double dy1 = (yup1-ylow1)/double(nby1); double dx2 = (xup2-xlow2)/double(nbx2); double dy2 = (yup2-ylow2)/double(nby2); // scale histo 2 to scale of 1 h2->Sumw2(); h2->Scale( ( double(n1) * dx1 * dy1 ) / ( double(n2) * dx2 * dy2 ) ); bool global = false; if (option > 10) global = true; if (global) { // fill data structure for fit (coordinates + values + errors) std::cout << "Do global fit" << std::endl; // fit now all the function together //The default minimizer is Minuit, you can also try Minuit2 if (option%10 == 2) TVirtualFitter::SetDefaultFitter("Minuit2"); else TVirtualFitter::SetDefaultFitter("Minuit"); TVirtualFitter * minuit = TVirtualFitter::Fitter(0,10); for (int i = 0; i < 10; ++i) { minuit->SetParameter(i, func->GetParName(i), func->GetParameter(i), 0.01, 0,0); } minuit->SetFCN(myFcn); double arglist[100]; arglist[0] = 0; // set print level minuit->ExecuteCommand("SET PRINT",arglist,2); // minimize arglist[0] = 5000; // number of function calls arglist[1] = 0.01; // tolerance minuit->ExecuteCommand("MIGRAD",arglist,2); //get result double minParams[10]; double parErrors[10]; for (int i = 0; i < 10; ++i) { minParams[i] = minuit->GetParameter(i); parErrors[i] = minuit->GetParError(i); } double chi2, edm, errdef; int nvpar, nparx; minuit->GetStats(chi2,edm,errdef,nvpar,nparx); func->SetParameters(minParams); func->SetParErrors(parErrors); func->SetChisquare(chi2); int ndf = npfits-nvpar; func->SetNDF(ndf); // add to list of functions h1->GetListOfFunctions()->Add(func); h2->GetListOfFunctions()->Add(func); } else { // fit independently h1->Fit(func); h2->Fit(func); } // Create a new canvas. TCanvas * c1 = new TCanvas("c1","Two HIstogram Fit example",100,10,900,800); c1->Divide(2,2); gStyle->SetOptFit(); gStyle->SetStatY(0.6); c1->cd(1); h1->Draw(); func->SetRange(xlow1,ylow1,xup1,yup1); func->DrawCopy("cont1 same"); c1->cd(2); h1->Draw("lego"); func->DrawCopy("surf1 same"); c1->cd(3); func->SetRange(xlow2,ylow2,xup2,yup2); h2->Draw(); func->DrawCopy("cont1 same"); c1->cd(4); h2->Draw("lego"); gPad->SetLogz(); func->Draw("surf1 same"); return 0; } fit2dHist.C:1 fit2dHist.C:2 fit2dHist.C:3 fit2dHist.C:4 fit2dHist.C:5 fit2dHist.C:6 fit2dHist.C:7 fit2dHist.C:8 fit2dHist.C:9 fit2dHist.C:10 fit2dHist.C:11 fit2dHist.C:12 fit2dHist.C:13 fit2dHist.C:14 fit2dHist.C:15 fit2dHist.C:16 fit2dHist.C:17 fit2dHist.C:18 fit2dHist.C:19 fit2dHist.C:20 fit2dHist.C:21 fit2dHist.C:22 fit2dHist.C:23 fit2dHist.C:24 fit2dHist.C:25 fit2dHist.C:26 fit2dHist.C:27 fit2dHist.C:28 fit2dHist.C:29 fit2dHist.C:30 fit2dHist.C:31 fit2dHist.C:32 fit2dHist.C:33 fit2dHist.C:34 fit2dHist.C:35 fit2dHist.C:36 fit2dHist.C:37 fit2dHist.C:38 fit2dHist.C:39 fit2dHist.C:40 fit2dHist.C:41 fit2dHist.C:42 fit2dHist.C:43 fit2dHist.C:44 fit2dHist.C:45 fit2dHist.C:46 fit2dHist.C:47 fit2dHist.C:48 fit2dHist.C:49 fit2dHist.C:50 fit2dHist.C:51 fit2dHist.C:52 fit2dHist.C:53 fit2dHist.C:54 fit2dHist.C:55 fit2dHist.C:56 fit2dHist.C:57 fit2dHist.C:58 fit2dHist.C:59 fit2dHist.C:60 fit2dHist.C:61 fit2dHist.C:62 fit2dHist.C:63 fit2dHist.C:64 fit2dHist.C:65 fit2dHist.C:66 fit2dHist.C:67 fit2dHist.C:68 fit2dHist.C:69 fit2dHist.C:70 fit2dHist.C:71 fit2dHist.C:72 fit2dHist.C:73 fit2dHist.C:74 fit2dHist.C:75 fit2dHist.C:76 fit2dHist.C:77 fit2dHist.C:78 fit2dHist.C:79 fit2dHist.C:80 fit2dHist.C:81 fit2dHist.C:82 fit2dHist.C:83 fit2dHist.C:84 fit2dHist.C:85 fit2dHist.C:86 fit2dHist.C:87 fit2dHist.C:88 fit2dHist.C:89 fit2dHist.C:90 fit2dHist.C:91 fit2dHist.C:92 fit2dHist.C:93 fit2dHist.C:94 fit2dHist.C:95 fit2dHist.C:96 fit2dHist.C:97 fit2dHist.C:98 fit2dHist.C:99 fit2dHist.C:100 fit2dHist.C:101 fit2dHist.C:102 fit2dHist.C:103 fit2dHist.C:104 fit2dHist.C:105 fit2dHist.C:106 fit2dHist.C:107 fit2dHist.C:108 fit2dHist.C:109 fit2dHist.C:110 fit2dHist.C:111 fit2dHist.C:112 fit2dHist.C:113 fit2dHist.C:114 fit2dHist.C:115 fit2dHist.C:116 fit2dHist.C:117 fit2dHist.C:118 fit2dHist.C:119 fit2dHist.C:120 fit2dHist.C:121 fit2dHist.C:122 fit2dHist.C:123 fit2dHist.C:124 fit2dHist.C:125 fit2dHist.C:126 fit2dHist.C:127 fit2dHist.C:128 fit2dHist.C:129 fit2dHist.C:130 fit2dHist.C:131 fit2dHist.C:132 fit2dHist.C:133 fit2dHist.C:134 fit2dHist.C:135 fit2dHist.C:136 fit2dHist.C:137 fit2dHist.C:138 fit2dHist.C:139 fit2dHist.C:140 fit2dHist.C:141 fit2dHist.C:142 fit2dHist.C:143 fit2dHist.C:144 fit2dHist.C:145 fit2dHist.C:146 fit2dHist.C:147 fit2dHist.C:148 fit2dHist.C:149 fit2dHist.C:150 fit2dHist.C:151 fit2dHist.C:152 fit2dHist.C:153 fit2dHist.C:154 fit2dHist.C:155 fit2dHist.C:156 fit2dHist.C:157 fit2dHist.C:158 fit2dHist.C:159 fit2dHist.C:160 fit2dHist.C:161 fit2dHist.C:162 fit2dHist.C:163 fit2dHist.C:164 fit2dHist.C:165 fit2dHist.C:166 fit2dHist.C:167 fit2dHist.C:168 fit2dHist.C:169 fit2dHist.C:170 fit2dHist.C:171 fit2dHist.C:172 fit2dHist.C:173 fit2dHist.C:174 fit2dHist.C:175 fit2dHist.C:176 fit2dHist.C:177 fit2dHist.C:178 fit2dHist.C:179 fit2dHist.C:180 fit2dHist.C:181 fit2dHist.C:182 fit2dHist.C:183 fit2dHist.C:184 fit2dHist.C:185 fit2dHist.C:186 fit2dHist.C:187 fit2dHist.C:188 fit2dHist.C:189 fit2dHist.C:190 fit2dHist.C:191 fit2dHist.C:192 fit2dHist.C:193 fit2dHist.C:194 fit2dHist.C:195 fit2dHist.C:196 fit2dHist.C:197 fit2dHist.C:198 fit2dHist.C:199 fit2dHist.C:200 fit2dHist.C:201 fit2dHist.C:202 fit2dHist.C:203 fit2dHist.C:204 fit2dHist.C:205 fit2dHist.C:206 fit2dHist.C:207 fit2dHist.C:208 fit2dHist.C:209 fit2dHist.C:210 fit2dHist.C:211 fit2dHist.C:212 fit2dHist.C:213 fit2dHist.C:214 fit2dHist.C:215 fit2dHist.C:216 fit2dHist.C:217 fit2dHist.C:218 fit2dHist.C:219 fit2dHist.C:220 fit2dHist.C:221 fit2dHist.C:222 fit2dHist.C:223 fit2dHist.C:224 fit2dHist.C:225 fit2dHist.C:226 fit2dHist.C:227 fit2dHist.C:228 fit2dHist.C:229 fit2dHist.C:230 fit2dHist.C:231 fit2dHist.C:232 fit2dHist.C:233 fit2dHist.C:234 fit2dHist.C:235 fit2dHist.C:236 fit2dHist.C:237 fit2dHist.C:238 fit2dHist.C:239 fit2dHist.C:240 fit2dHist.C:241 fit2dHist.C:242 |
|