| TwoSidedFrequentistUpperLimitWithBands.C: | Roostats tutorials | rs101_limitexample.C: 'Limit Example' RooStats tutorial macro #101 |
Zbi_Zgamma.C: Demonstraite Z_Bi = Z_Gamma
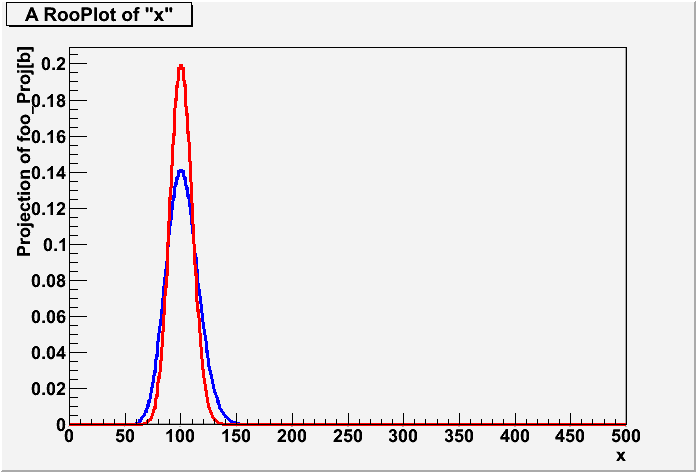
///////////////////////////////////////////////////////////////////////// // // Demonstraite Z_Bi = Z_Gamma // author: Kyle Cranmer & Wouter Verkerke // date May 2010 // // ///////////////////////////////////////////////////////////////////////// #ifndef __CINT__ #include "RooGlobalFunc.h" #endif #include "RooRealVar.h" #include "RooProdPdf.h" #include "RooWorkspace.h" #include "RooDataSet.h" #include "TCanvas.h" #include "TH1.h" using namespace RooFit; using namespace RooStats; void Zbi_Zgamma() { // Make model for prototype on/off problem // Pois(x | s+b) * Pois(y | tau b ) // for Z_Gamma, use uniform prior on b. RooWorkspace* w = new RooWorkspace("w",true); w->factory("Poisson::px(x[150,0,500],sum::splusb(s[0,0,100],b[100,0,300]))"); w->factory("Poisson::py(y[100,0,500],prod::taub(tau[1.],b))"); w->factory("Uniform::prior_b(b)"); // construct the Bayesian-averaged model (eg. a projection pdf) // p'(x|s) = \int db p(x|s+b) * [ p(y|b) * prior(b) ] w->factory("PROJ::averagedModel(PROD::foo(px|b,py,prior_b),b)") ; // plot it, blue is averaged model, red is b known exactly RooPlot* frame = w->var("x")->frame() ; w->pdf("averagedModel")->plotOn(frame) ; w->pdf("px")->plotOn(frame,LineColor(kRed)) ; frame->Draw() ; // compare analytic calculation of Z_Bi // with the numerical RooFit implementation of Z_Gamma // for an example with x = 150, y = 100 // numeric RooFit Z_Gamma w->var("y")->setVal(100); w->var("x")->setVal(150); RooAbsReal* cdf = w->pdf("averagedModel")->createCdf(*w->var("x")); cdf->getVal(); // get ugly print messages out of the way cout << "Hybrid p-value = " << cdf->getVal() << endl; cout << "Z_Gamma Significance = " << PValueToSignificance(1-cdf->getVal()) << endl; // analytic Z_Bi double Z_Bi = NumberCountingUtils::BinomialWithTauObsZ(150, 100, 1); std::cout << "Z_Bi significance estimation: " << Z_Bi << std::endl; // OUTPUT // Hybrid p-value = 0.999058 // Z_Gamma Significance = 3.10804 // Z_Bi significance estimation: 3.10804 } Zbi_Zgamma.C:1 Zbi_Zgamma.C:2 Zbi_Zgamma.C:3 Zbi_Zgamma.C:4 Zbi_Zgamma.C:5 Zbi_Zgamma.C:6 Zbi_Zgamma.C:7 Zbi_Zgamma.C:8 Zbi_Zgamma.C:9 Zbi_Zgamma.C:10 Zbi_Zgamma.C:11 Zbi_Zgamma.C:12 Zbi_Zgamma.C:13 Zbi_Zgamma.C:14 Zbi_Zgamma.C:15 Zbi_Zgamma.C:16 Zbi_Zgamma.C:17 Zbi_Zgamma.C:18 Zbi_Zgamma.C:19 Zbi_Zgamma.C:20 Zbi_Zgamma.C:21 Zbi_Zgamma.C:22 Zbi_Zgamma.C:23 Zbi_Zgamma.C:24 Zbi_Zgamma.C:25 Zbi_Zgamma.C:26 Zbi_Zgamma.C:27 Zbi_Zgamma.C:28 Zbi_Zgamma.C:29 Zbi_Zgamma.C:30 Zbi_Zgamma.C:31 Zbi_Zgamma.C:32 Zbi_Zgamma.C:33 Zbi_Zgamma.C:34 Zbi_Zgamma.C:35 Zbi_Zgamma.C:36 Zbi_Zgamma.C:37 Zbi_Zgamma.C:38 Zbi_Zgamma.C:39 Zbi_Zgamma.C:40 Zbi_Zgamma.C:41 Zbi_Zgamma.C:42 Zbi_Zgamma.C:43 Zbi_Zgamma.C:44 Zbi_Zgamma.C:45 Zbi_Zgamma.C:46 Zbi_Zgamma.C:47 Zbi_Zgamma.C:48 Zbi_Zgamma.C:49 Zbi_Zgamma.C:50 Zbi_Zgamma.C:51 Zbi_Zgamma.C:52 Zbi_Zgamma.C:53 Zbi_Zgamma.C:54 Zbi_Zgamma.C:55 Zbi_Zgamma.C:56 Zbi_Zgamma.C:57 Zbi_Zgamma.C:58 Zbi_Zgamma.C:59 Zbi_Zgamma.C:60 Zbi_Zgamma.C:61 Zbi_Zgamma.C:62 Zbi_Zgamma.C:63 Zbi_Zgamma.C:64 Zbi_Zgamma.C:65 Zbi_Zgamma.C:66 Zbi_Zgamma.C:67 Zbi_Zgamma.C:68 |
|