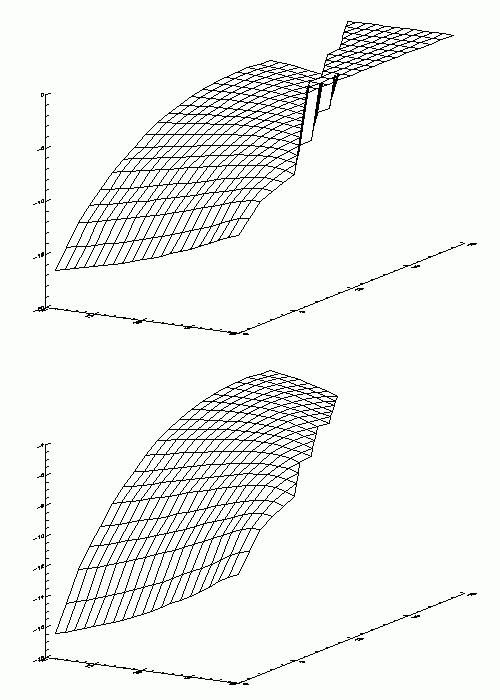
Sorry, I'll try to be a little more clear. What actually happens when you designate a range of values or individual values as "bad" or "missing" is that they are then ignored by the plotting routine. An example plotted using IDL (but generated using by dumping a TProfile2D class after analysis to an ASCII file) is given below. The upper diagram corresponds to "no missing values" , whereas for the lower graph I've simply specified that all values above -1 dB are "missing" data and should not by plotted. Similar results can be obtained for the IDL contour routines. The missing data in this case are simply histogram bins which have not been filled. Other examples include NaNs which one does not want to plot etc...One can get around this behaviour to a certain exent by setting "missing" histogram bins to some sensible value not too far from the mean of the non-zero bins but this still doesn't quite give the same result. Hope this helps in clarifying things. I realise that changing the plotting routines to include this kind of behaviour is relatively onerous but it is only a wish...one which I guess other users might have as well. Regards Malcolm At 21:55 11/04/01 +0200, Rene Brun wrote: >Hi Malcolm, > >I am certainly missing some elements in your request. >It seems to be more logical to eliminate the bad values while you >fill your histogram than a posteriori. >With Root, you typically store your data as an ntuple (Tree) and project >the data to an histogram with a selection. > >Rene Brun > >On Wed, 11 Apr 2001, Malcolm Davidson wrote: > > > Hello Root Developers, > > > > I don't know if "missing value" support in displaying histogram and graph > > plots has already been added to the wish list ? Does anybody else miss > this > > feature ? > > > > It's one of the features that I really like in IDL and am currently sorely > > missing in my ROOT analysis. It consists basically in the the ability to > > designate certain values as "missing" or "bad" data and have the > > contour/plot routines ignore this data when building up the > > image/contour/plot. > > > > Continue the good work. > > > > Malcolm > > > > <>------------------------------------------------<> > > Malcolm W. J. Davidson > > CESBIO - UMR 5639 CNES-CNRS-UPS > > 18 Avenue Edouard Belin > > BP 2801 > > F-31401 Toulouse Cedex 4 France > > e-mail : davidson@cesbio.cnes.fr > > phone (33)(0)5.61.55.85.84 > > fax (33)(0)5.61.55.85.00 > > <>------------------------------------------------<> > >
 <>------------------------------------------------<>
Malcolm W. J. Davidson
CESBIO - UMR 5639 CNES-CNRS-UPS
18 Avenue Edouard Belin
BP 2801
F-31401 Toulouse Cedex 4 France
e-mail : davidson@cesbio.cnes.fr
phone (33)(0)5.61.55.85.84
fax (33)(0)5.61.55.85.00
<>------------------------------------------------<>
<>------------------------------------------------<>
Malcolm W. J. Davidson
CESBIO - UMR 5639 CNES-CNRS-UPS
18 Avenue Edouard Belin
BP 2801
F-31401 Toulouse Cedex 4 France
e-mail : davidson@cesbio.cnes.fr
phone (33)(0)5.61.55.85.84
fax (33)(0)5.61.55.85.00
<>------------------------------------------------<>
This archive was generated by hypermail 2b29 : Tue Jan 01 2002 - 17:50:41 MET