

Comparison of MCMC and PLC in a multi-variate gaussian problem
This tutorial produces an N-dimensional multivariate Gaussian with a non-trivial covariance matrix. By default N=4 (called "dim").
A subset of these are considered parameters of interest. This problem is tractable analytically.
We use this mainly as a test of Markov Chain Monte Carlo and we compare the result to the profile likelihood ratio.
We use the proposal helper to create a customized proposal function for this problem.
For N=4 and 2 parameters of interest it takes about 10-20 seconds and the acceptance rate is 37%
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
Minuit2Minimizer: Minimize with max-calls 2000 convergence for edm < 1 strategy 1
Minuit2Minimizer : Valid minimum - status = 0
FVAL = 706.560063684865781
Edm = 9.79361278577427602e-06
Nfcn = 68
mu_x0 = 0.180728 +/- 0.17298 (limited)
mu_x1 = 0.207351 +/- 0.172978 (limited)
mu_x2 = -0.0159412 +/- 0.172984 (limited)
mu_x3 = 0.12343 +/- 0.172982 (limited)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
Metropolis-Hastings progress: ....................................................................................................
[#1] INFO:Eval -- Proposal acceptance rate: 37.1%
[#1] INFO:Eval -- Number of steps in chain: 3710
[#1] INFO:InputArguments -- The deprecated RooFit::CloneData(1) option passed to createNLL() is ignored.
[#0] PROGRESS:Minimization -- ProfileLikelihoodCalcultor::DoGLobalFit - find MLE
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#0] PROGRESS:Minimization -- ProfileLikelihoodCalcultor::DoMinimizeNLL - using Minuit2 / Migrad with strategy 1
[#1] INFO:Minimization --
RooFitResult: minimized FCN value: 706.56, estimated distance to minimum: 3.96149e-12
covariance matrix quality: Full, accurate covariance matrix
Status : MINIMIZE=0
Floating Parameter FinalValue +/- Error
-------------------- --------------------------
mu_x0 1.8118e-01 +/- 1.73e-01
mu_x1 2.0792e-01 +/- 1.73e-01
mu_x2 -1.6067e-02 +/- 1.73e-01
mu_x3 1.2371e-01 +/- 1.73e-01
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_mvg_mvgData_Profile[mu_x1,mu_x0]) Creating instance of MINUIT
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_mvg_mvgData_Profile[mu_x1,mu_x0]) determining minimum likelihood for current configurations w.r.t all observable
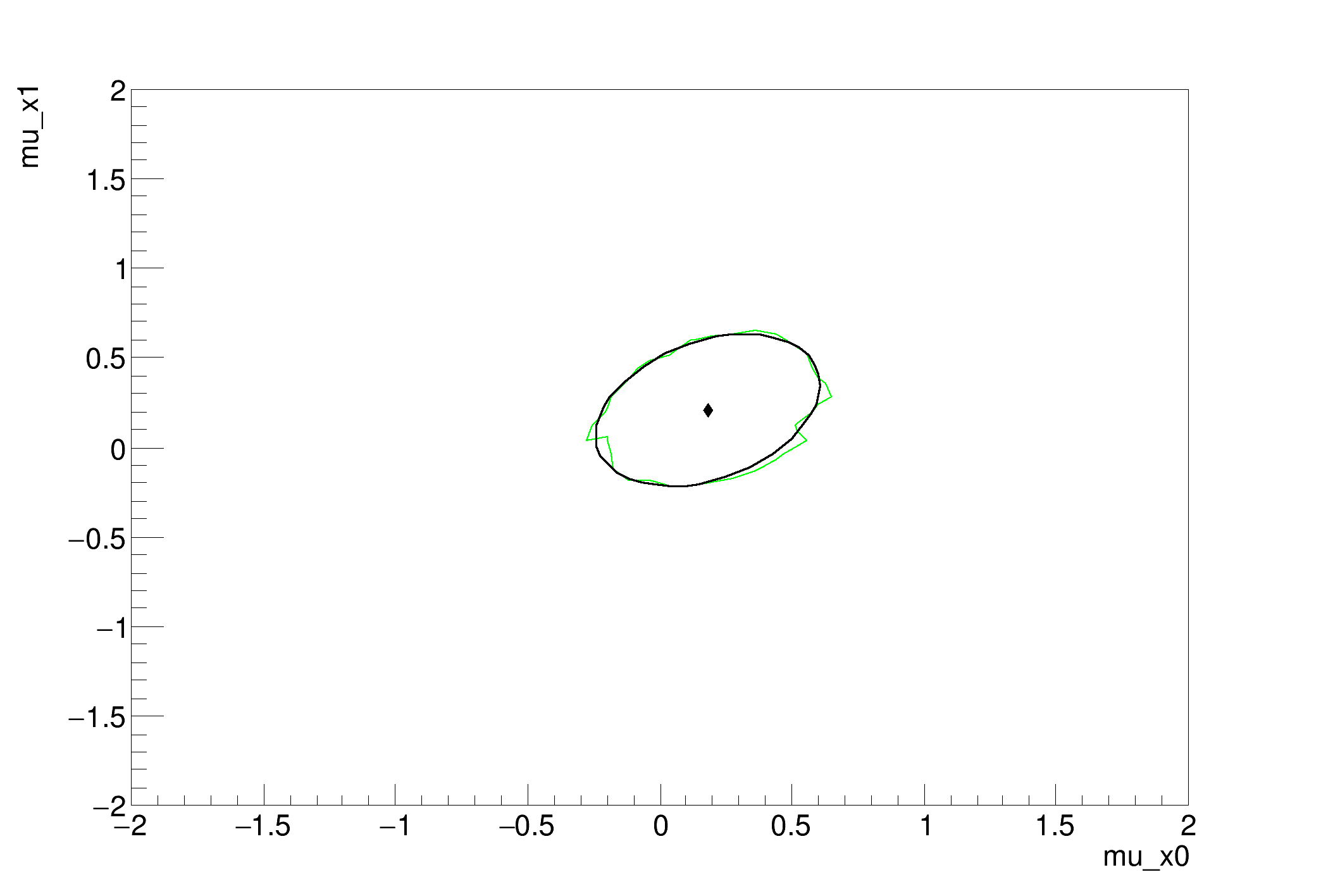
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_mvg_mvgData_Profile[mu_x1,mu_x0]) minimum found at (mu_x0=0.181184, mu_x1=0.207918)
..[#1] INFO:Minimization -- LikelihoodInterval - Finding the contour of mu_x1 ( 1 ) and mu_x0 ( 0 )
Real time 0:00:02, CP time 2.950
MCMC interval on p0: [-0.19999999999999996, 0.6000000000000001]
MCMC interval on p1: [-0.28, 0.6000000000000001]
import ROOT
dim = 4
nPOI = 2
t = ROOT.TStopwatch()
t.Start()
xVec = []
muVec = []
poi = set()
for i in range(dim):
x = ROOT.RooRealVar(name, name, 0, -3, 3)
xVec.append(x)
mu_x = ROOT.RooRealVar(mu_name, mu_name, 0, -2, 2)
muVec.append(mu_x)
for i in range(nPOI):
poi.add(muVec[i])
cov = ROOT.TMatrixDSym(dim)
for i in range(dim):
for j in range(dim):
if i == j:
cov[i, j] = 3.0
else:
cov[i, j] = 1.0
mvg = ROOT.RooMultiVarGaussian("mvg", "mvg", xVec, muVec, cov)
data = mvg.generate(xVec, 100)
w = ROOT.RooWorkspace("MVG")
modelConfig = ROOT.RooStats.ModelConfig(w)
modelConfig.SetPdf(mvg)
modelConfig.SetParametersOfInterest(poi)
fit = mvg.fitTo(data, Save=True)
ph = ROOT.RooStats.ProposalHelper()
ph.SetVariables(fit.floatParsFinal())
ph.SetCovMatrix(fit.covarianceMatrix())
ph.SetUpdateProposalParameters(True)
ph.SetCacheSize(100)
pdfProp = ph.GetProposalFunction()
mc = ROOT.RooStats.MCMCCalculator(data, modelConfig)
mc.SetConfidenceLevel(0.95)
mc.SetNumBurnInSteps(100)
mc.SetNumIters(10000)
mc.SetNumBins(50)
mc.SetProposalFunction(pdfProp)
mcInt = mc.GetInterval()
poiList = mcInt.GetAxes()
plc = ROOT.RooStats.ProfileLikelihoodCalculator(data, modelConfig)
plc.SetConfidenceLevel(0.95)
plInt = plc.GetInterval()
mcPlot = ROOT.RooStats.MCMCIntervalPlot(mcInt)
c1 = ROOT.TCanvas()
mcPlot.SetLineColor(ROOT.kGreen)
mcPlot.SetLineWidth(2)
mcPlot.Draw()
plPlot = ROOT.RooStats.LikelihoodIntervalPlot(plInt)
plPlot.Draw("same")
p = poiList.at(0)
print(
"MCMC interval: [{}, {}]".
format(mcInt.LowerLimit(p), mcInt.UpperLimit(p)))
p0 = poiList.at(0)
p1 = poiList.at(1)
scatter = ROOT.TCanvas()
print(
"MCMC interval on p0: [{}, {}]".
format(mcInt.LowerLimit(p0), mcInt.UpperLimit(p0)))
print(
"MCMC interval on p1: [{}, {}]".
format(mcInt.LowerLimit(p1), mcInt.UpperLimit(p1)))
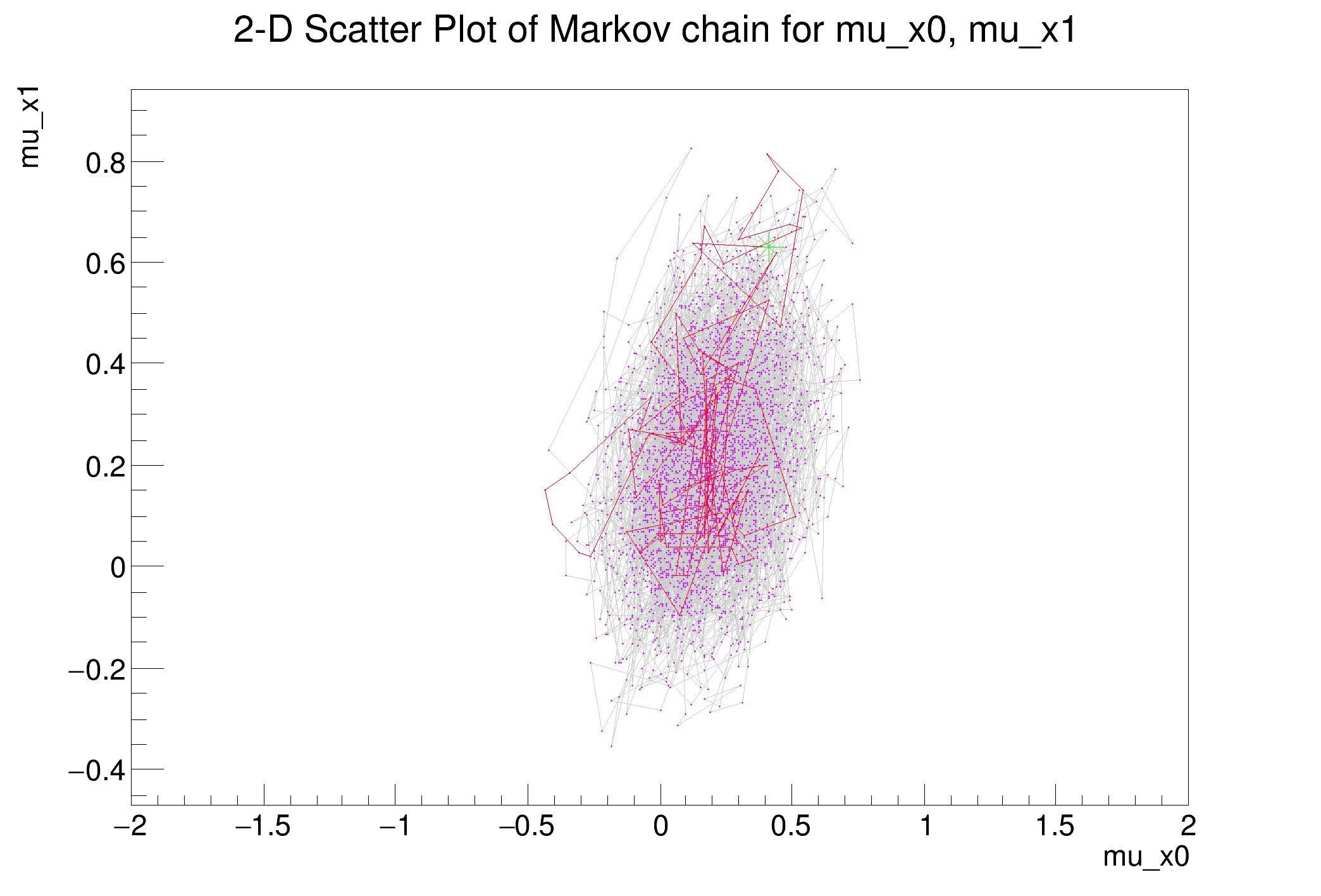
mcPlot.DrawChainScatter(p0, p1)
scatter.Update()
scatter.SaveAs("MultivariateGaussianTest_scatter.png")
t.Stop()
t.Print()
c1.SaveAs("MultivariateGaussianTest_plot.png")
del mc
Option_t Option_t TPoint TPoint const char GetTextMagnitude GetFillStyle GetLineColor GetLineWidth GetMarkerStyle GetTextAlign GetTextColor GetTextSize void char Point_t Rectangle_t WindowAttributes_t Float_t Float_t Float_t Int_t Int_t UInt_t UInt_t Rectangle_t Int_t Int_t Window_t TString Int_t GCValues_t GetPrimarySelectionOwner GetDisplay GetScreen GetColormap GetNativeEvent const char const char dpyName wid window const char font_name cursor keysym reg const char only_if_exist regb h Point_t winding char text const char depth char const char Int_t count const char ColorStruct_t color const char Pixmap_t Pixmap_t PictureAttributes_t attr const char char ret_data h unsigned char height h Atom_t Int_t ULong_t ULong_t unsigned char prop_list Atom_t Atom_t Atom_t Time_t UChar_t len
Option_t Option_t TPoint TPoint const char GetTextMagnitude GetFillStyle GetLineColor GetLineWidth GetMarkerStyle GetTextAlign GetTextColor GetTextSize void char Point_t Rectangle_t WindowAttributes_t Float_t Float_t Float_t Int_t Int_t UInt_t UInt_t Rectangle_t Int_t Int_t Window_t TString Int_t GCValues_t GetPrimarySelectionOwner GetDisplay GetScreen GetColormap GetNativeEvent const char const char dpyName wid window const char font_name cursor keysym reg const char only_if_exist regb h Point_t winding char text const char depth char const char Int_t count const char ColorStruct_t color const char Pixmap_t Pixmap_t PictureAttributes_t attr const char char ret_data h unsigned char height h Atom_t Int_t ULong_t ULong_t unsigned char prop_list Atom_t Atom_t Atom_t Time_t format
- Date
- July 2022
- Authors
- Artem Busorgin, Kevin Belasco and Kyle Cranmer (C++ version)
Definition in file MultivariateGaussianTest.py.



