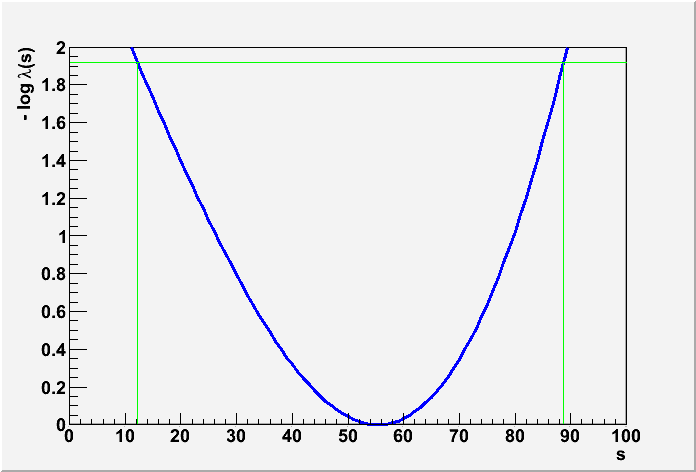
FourBinInstructional.C: This example is a generalization of the on/off problem.
// This example is a generalization of the on/off problem. /* FourBin Instructional Tutorial: authors: Kyle Cranmer <cranmer@cern.ch> Tanja Rommerskirchen <tanja.rommerskirchen@cern.ch> date: June 1, 2010 This example is a generalization of the on/off problem. It's a common setup for SUSY searches. Imagine that one has two variables "x" and "y" (eg. missing ET and SumET), see figure. The signal region has high values of both of these variables (top right). One can see low values of "x" or "y" acting as side-bands. If we just used "y" as a sideband, we would have the on/off problem. - In the signal region we observe non events and expect s+b events. - In the region with low values of "y" (bottom right) we observe noff events and expect tau*b events. Note the significance of tau. In the background only case: tau ~ <expectation off> / <expectation on> If tau is known, this model is sufficient, but often tau is not known exactly. So one can use low values of "x" as an additional constraint for tau. Note that this technique critically depends on the notion that the joint distribution for "x" and "y" can be factorized. Generally, these regions have many events, so it the ratio can be measured very precisely there. So we extend the model to describe the left two boxes... denoted with "bar". - In the upper left we observe nonbar events and expect bbar events - In the bottom left we observe noffbar events and expect tau bbar events Note again we have: tau ~ <expecation off bar> / <expectation on bar> One can further expand the model to account for the systematic associated to assuming the distribution of "x" and "y" factorizes (eg. that tau is the same for off/on and offbar/onbar). This can be done in several ways, but here we introduce an additional parameter rho, which so that one set of models will use tau and the other tau*rho. The choice is arbitary, but it has consequences on the numerical stability of the algorithms. The "bar" measurements typically have more events (& smaller relative errors). If we choose <expectation noffbar> = tau * rho * <expectation noonbar>, the product tau*rho will be known very precisely (~1/sqrt(bbar)) and the contour in those parameters will be narrow and have a non-trivial tau~1/rho shape. However, if we choose to put rho on the non/noff measurements (where the product will have an error ~1/sqrt(b)), the contours will be more ameanable to numerical techniques. Thus, here we choose to define tau := <expecation off bar> / (<expectation on bar>) rho := <expecation off> / (<expectation on> * tau) ^ y | |---------------------------+ | | | | nonbar | non | | bbar | s+b | | | | |---------------+-----------| | | | | noffbar | noff | | tau bbar | tau b rho | | | | +-----------------------------> x Left in this way, the problem is under-constrained. However, one may have some auxiliary measurement (usually based on Monte Carlo) to constrain rho. Let us call this auxiliary measurement that gives the nominal value of rho "rhonom". Thus, there is a 'constraint' term in the full model: P(rhonom | rho). In this case, we consider a Gaussian constraint with standard deviation sigma. In the example, the initial values of the parameters are - s = 40 - b = 100 - tau = 5 - bbar = 1000 - rho = 1 (sigma for rho = 20%) and in the toy dataset: - non = 139 - noff = 528 - nonbar = 993 - noffbar = 4906 - rhonom = 1.27824 Note, the covariance matrix of the parameters has large off-diagonal terms. Clearly s,b are anti-correlated. Similary, since noffbar >> nonbar, one would expect bbar,tau to be anti-correlated. This can be seen below. GLOBAL b bbar rho s tau b 0.96820 1.000 0.191 -0.942 -0.762 -0.209 bbar 0.91191 0.191 1.000 0.000 -0.146 -0.912 rho 0.96348 -0.942 0.000 1.000 0.718 -0.000 s 0.76250 -0.762 -0.146 0.718 1.000 0.160 tau 0.92084 -0.209 -0.912 -0.000 0.160 1.000 Similarly, since tau*rho appears as a product, we expect rho,tau to be anti-correlated. When the error on rho is significantly larger than 1/sqrt(bbar), tau is essentially known and the correlation is minimal (tau mainly cares about bbar, and rho about b,s). In the alternate parametrizaton (bbar* tau * rho) the correlation coefficient for rho,tau is large (and negative). The code below uses best-practices for RooFit & RooStats as of June 2010. It proceeds as follows: - create a workspace to hold the model - use workspace factory to quickly create the terms of the model - use workspace factory to define total model (a prod pdf) - create a RooStats ModelConfig to specify observables, parameters of interest - add to the ModelConfig a prior on the parameters for Bayesian techniques - note, the pdf it is factorized for parameters of interest & nuisance params - visualize the model - write the workspace to a file - use several of RooStats IntervalCalculators & compare results */ #include "TStopwatch.h" #include "TCanvas.h" #include "TROOT.h" #include "RooPlot.h" #include "RooAbsPdf.h" #include "RooWorkspace.h" #include "RooDataSet.h" #include "RooGlobalFunc.h" #include "RooFitResult.h" #include "RooRandom.h" #include "RooStats/ProfileLikelihoodCalculator.h" #include "RooStats/LikelihoodInterval.h" #include "RooStats/LikelihoodIntervalPlot.h" #include "RooStats/BayesianCalculator.h" #include "RooStats/MCMCCalculator.h" #include "RooStats/MCMCInterval.h" #include "RooStats/MCMCIntervalPlot.h" #include "RooStats/ProposalHelper.h" #include "RooStats/SimpleInterval.h" #include "RooStats/FeldmanCousins.h" #include "RooStats/PointSetInterval.h" using namespace RooFit; using namespace RooStats; void FourBinInstructional(bool doBayesian=false, bool doFeldmanCousins=false, bool doMCMC=false){ // let's time this challenging example TStopwatch t; t.Start(); // set RooFit random seed for reproducible results RooRandom::randomGenerator()->SetSeed(4357); // make model RooWorkspace* wspace = new RooWorkspace("wspace"); wspace->factory("Poisson::on(non[0,1000], sum::splusb(s[40,0,100],b[100,0,300]))"); wspace->factory("Poisson::off(noff[0,5000], prod::taub(b,tau[5,3,7],rho[1,0,2]))"); wspace->factory("Poisson::onbar(nonbar[0,10000], bbar[1000,500,2000])"); wspace->factory("Poisson::offbar(noffbar[0,1000000], prod::lambdaoffbar(bbar, tau))"); wspace->factory("Gaussian::mcCons(rhonom[1.,0,2], rho, sigma[.2])"); wspace->factory("PROD::model(on,off,onbar,offbar,mcCons)"); wspace->defineSet("obs","non,noff,nonbar,noffbar,rhonom"); wspace->factory("Uniform::prior_poi({s})"); wspace->factory("Uniform::prior_nuis({b,bbar,tau, rho})"); wspace->factory("PROD::prior(prior_poi,prior_nuis)"); /////////////////////////////////////////// // Control some interesting variations // define parameers of interest // for 1-d plots wspace->defineSet("poi","s"); wspace->defineSet("nuis","b,tau,rho,bbar"); // for 2-d plots to inspect correlations: // wspace->defineSet("poi","s,rho"); // test simpler cases where parameters are known. // wspace->var("tau")->setConstant(); // wspace->var("rho")->setConstant(); // wspace->var("b")->setConstant(); // wspace->var("bbar")->setConstant(); // inspect workspace // wspace->Print(); //////////////////////////////////////////////////////////// // Generate toy data // generate toy data assuming current value of the parameters // import into workspace. // add Verbose() to see how it's being generated RooDataSet* data = wspace->pdf("model")->generate(*wspace->set("obs"),1); // data->Print("v"); wspace->import(*data); ///////////////////////////////////////////////////// // Now the statistical tests // model config ModelConfig* modelConfig = new ModelConfig("FourBins"); modelConfig->SetWorkspace(*wspace); modelConfig->SetPdf(*wspace->pdf("model")); modelConfig->SetPriorPdf(*wspace->pdf("prior")); modelConfig->SetParametersOfInterest(*wspace->set("poi")); modelConfig->SetNuisanceParameters(*wspace->set("nuis")); wspace->import(*modelConfig); wspace->writeToFile("FourBin.root"); ////////////////////////////////////////////////// // If you want to see the covariance matrix uncomment // wspace->pdf("model")->fitTo(*data); // use ProfileLikelihood ProfileLikelihoodCalculator plc(*data, *modelConfig); plc.SetConfidenceLevel(0.95); LikelihoodInterval* plInt = plc.GetInterval(); RooFit::MsgLevel msglevel = RooMsgService::instance().globalKillBelow(); RooMsgService::instance().setGlobalKillBelow(RooFit::FATAL); plInt->LowerLimit( *wspace->var("s") ); // get ugly print out of the way. Fix. RooMsgService::instance().setGlobalKillBelow(msglevel); // use FeldmaCousins (takes ~20 min) FeldmanCousins fc(*data, *modelConfig); fc.SetConfidenceLevel(0.95); //number counting: dataset always has 1 entry with N events observed fc.FluctuateNumDataEntries(false); fc.UseAdaptiveSampling(true); fc.SetNBins(40); PointSetInterval* fcInt = NULL; if(doFeldmanCousins){ // takes 7 minutes fcInt = (PointSetInterval*) fc.GetInterval(); // fix cast } // use BayesianCalculator (only 1-d parameter of interest, slow for this problem) BayesianCalculator bc(*data, *modelConfig); bc.SetConfidenceLevel(0.95); SimpleInterval* bInt = NULL; if(doBayesian && wspace->set("poi")->getSize() == 1) { bInt = bc.GetInterval(); } else{ cout << "Bayesian Calc. only supports on parameter of interest" << endl; } // use MCMCCalculator (takes about 1 min) // Want an efficient proposal function, so derive it from covariance // matrix of fit RooFitResult* fit = wspace->pdf("model")->fitTo(*data,Save()); ProposalHelper ph; ph.SetVariables((RooArgSet&)fit->floatParsFinal()); ph.SetCovMatrix(fit->covarianceMatrix()); ph.SetUpdateProposalParameters(kTRUE); // auto-create mean vars and add mappings ph.SetCacheSize(100); ProposalFunction* pf = ph.GetProposalFunction(); MCMCCalculator mc(*data, *modelConfig); mc.SetConfidenceLevel(0.95); mc.SetProposalFunction(*pf); mc.SetNumBurnInSteps(500); // first N steps to be ignored as burn-in mc.SetNumIters(50000); mc.SetLeftSideTailFraction(0.5); // make a central interval MCMCInterval* mcInt = NULL; if(doMCMC) mcInt = mc.GetInterval(); ////////////////////////////////////// // Make some plots TCanvas* c1 = (TCanvas*) gROOT->Get("c1"); if(!c1) c1 = new TCanvas("c1"); if(doBayesian && doMCMC){ c1->Divide(3); c1->cd(1); } else if (doBayesian || doMCMC){ c1->Divide(2); c1->cd(1); } LikelihoodIntervalPlot* lrplot = new LikelihoodIntervalPlot(plInt); lrplot->Draw(); if(doBayesian && wspace->set("poi")->getSize() == 1) { c1->cd(2); // the plot takes a long time and print lots of error // using a scan it is better bc.SetScanOfPosterior(20); RooPlot* bplot = bc.GetPosteriorPlot(); bplot->Draw(); } if(doMCMC){ if(doBayesian && wspace->set("poi")->getSize() == 1) c1->cd(3); else c1->cd(2); MCMCIntervalPlot mcPlot(*mcInt); mcPlot.Draw(); } //////////////////////////////////// // querry intervals cout << "Profile Likelihood interval on s = [" << plInt->LowerLimit( *wspace->var("s") ) << ", " << plInt->UpperLimit( *wspace->var("s") ) << "]" << endl; //Profile Likelihood interval on s = [12.1902, 88.6871] if(doBayesian && wspace->set("poi")->getSize() == 1) { cout << "Bayesian interval on s = [" << bInt->LowerLimit( ) << ", " << bInt->UpperLimit( ) << "]" << endl; } if(doFeldmanCousins){ cout << "Feldman Cousins interval on s = [" << fcInt->LowerLimit( *wspace->var("s") ) << ", " << fcInt->UpperLimit( *wspace->var("s") ) << "]" << endl; //Feldman Cousins interval on s = [18.75 +/- 2.45, 83.75 +/- 2.45] } if(doMCMC){ cout << "MCMC interval on s = [" << mcInt->LowerLimit(*wspace->var("s") ) << ", " << mcInt->UpperLimit(*wspace->var("s") ) << "]" << endl; //MCMC interval on s = [15.7628, 84.7266] } t.Print(); } FourBinInstructional.C:1 FourBinInstructional.C:2 FourBinInstructional.C:3 FourBinInstructional.C:4 FourBinInstructional.C:5 FourBinInstructional.C:6 FourBinInstructional.C:7 FourBinInstructional.C:8 FourBinInstructional.C:9 FourBinInstructional.C:10 FourBinInstructional.C:11 FourBinInstructional.C:12 FourBinInstructional.C:13 FourBinInstructional.C:14 FourBinInstructional.C:15 FourBinInstructional.C:16 FourBinInstructional.C:17 FourBinInstructional.C:18 FourBinInstructional.C:19 FourBinInstructional.C:20 FourBinInstructional.C:21 FourBinInstructional.C:22 FourBinInstructional.C:23 FourBinInstructional.C:24 FourBinInstructional.C:25 FourBinInstructional.C:26 FourBinInstructional.C:27 FourBinInstructional.C:28 FourBinInstructional.C:29 FourBinInstructional.C:30 FourBinInstructional.C:31 FourBinInstructional.C:32 FourBinInstructional.C:33 FourBinInstructional.C:34 FourBinInstructional.C:35 FourBinInstructional.C:36 FourBinInstructional.C:37 FourBinInstructional.C:38 FourBinInstructional.C:39 FourBinInstructional.C:40 FourBinInstructional.C:41 FourBinInstructional.C:42 FourBinInstructional.C:43 FourBinInstructional.C:44 FourBinInstructional.C:45 FourBinInstructional.C:46 FourBinInstructional.C:47 FourBinInstructional.C:48 FourBinInstructional.C:49 FourBinInstructional.C:50 FourBinInstructional.C:51 FourBinInstructional.C:52 FourBinInstructional.C:53 FourBinInstructional.C:54 FourBinInstructional.C:55 FourBinInstructional.C:56 FourBinInstructional.C:57 FourBinInstructional.C:58 FourBinInstructional.C:59 FourBinInstructional.C:60 FourBinInstructional.C:61 FourBinInstructional.C:62 FourBinInstructional.C:63 FourBinInstructional.C:64 FourBinInstructional.C:65 FourBinInstructional.C:66 FourBinInstructional.C:67 FourBinInstructional.C:68 FourBinInstructional.C:69 FourBinInstructional.C:70 FourBinInstructional.C:71 FourBinInstructional.C:72 FourBinInstructional.C:73 FourBinInstructional.C:74 FourBinInstructional.C:75 FourBinInstructional.C:76 FourBinInstructional.C:77 FourBinInstructional.C:78 FourBinInstructional.C:79 FourBinInstructional.C:80 FourBinInstructional.C:81 FourBinInstructional.C:82 FourBinInstructional.C:83 FourBinInstructional.C:84 FourBinInstructional.C:85 FourBinInstructional.C:86 FourBinInstructional.C:87 FourBinInstructional.C:88 FourBinInstructional.C:89 FourBinInstructional.C:90 FourBinInstructional.C:91 FourBinInstructional.C:92 FourBinInstructional.C:93 FourBinInstructional.C:94 FourBinInstructional.C:95 FourBinInstructional.C:96 FourBinInstructional.C:97 FourBinInstructional.C:98 FourBinInstructional.C:99 FourBinInstructional.C:100 FourBinInstructional.C:101 FourBinInstructional.C:102 FourBinInstructional.C:103 FourBinInstructional.C:104 FourBinInstructional.C:105 FourBinInstructional.C:106 FourBinInstructional.C:107 FourBinInstructional.C:108 FourBinInstructional.C:109 FourBinInstructional.C:110 FourBinInstructional.C:111 FourBinInstructional.C:112 FourBinInstructional.C:113 FourBinInstructional.C:114 FourBinInstructional.C:115 FourBinInstructional.C:116 FourBinInstructional.C:117 FourBinInstructional.C:118 FourBinInstructional.C:119 FourBinInstructional.C:120 FourBinInstructional.C:121 FourBinInstructional.C:122 FourBinInstructional.C:123 FourBinInstructional.C:124 FourBinInstructional.C:125 FourBinInstructional.C:126 FourBinInstructional.C:127 FourBinInstructional.C:128 FourBinInstructional.C:129 FourBinInstructional.C:130 FourBinInstructional.C:131 FourBinInstructional.C:132 FourBinInstructional.C:133 FourBinInstructional.C:134 FourBinInstructional.C:135 FourBinInstructional.C:136 FourBinInstructional.C:137 FourBinInstructional.C:138 FourBinInstructional.C:139 FourBinInstructional.C:140 FourBinInstructional.C:141 FourBinInstructional.C:142 FourBinInstructional.C:143 FourBinInstructional.C:144 FourBinInstructional.C:145 FourBinInstructional.C:146 FourBinInstructional.C:147 FourBinInstructional.C:148 FourBinInstructional.C:149 FourBinInstructional.C:150 FourBinInstructional.C:151 FourBinInstructional.C:152 FourBinInstructional.C:153 FourBinInstructional.C:154 FourBinInstructional.C:155 FourBinInstructional.C:156 FourBinInstructional.C:157 FourBinInstructional.C:158 FourBinInstructional.C:159 FourBinInstructional.C:160 FourBinInstructional.C:161 FourBinInstructional.C:162 FourBinInstructional.C:163 FourBinInstructional.C:164 FourBinInstructional.C:165 FourBinInstructional.C:166 FourBinInstructional.C:167 FourBinInstructional.C:168 FourBinInstructional.C:169 FourBinInstructional.C:170 FourBinInstructional.C:171 FourBinInstructional.C:172 FourBinInstructional.C:173 FourBinInstructional.C:174 FourBinInstructional.C:175 FourBinInstructional.C:176 FourBinInstructional.C:177 FourBinInstructional.C:178 FourBinInstructional.C:179 FourBinInstructional.C:180 FourBinInstructional.C:181 FourBinInstructional.C:182 FourBinInstructional.C:183 FourBinInstructional.C:184 FourBinInstructional.C:185 FourBinInstructional.C:186 FourBinInstructional.C:187 FourBinInstructional.C:188 FourBinInstructional.C:189 FourBinInstructional.C:190 FourBinInstructional.C:191 FourBinInstructional.C:192 FourBinInstructional.C:193 FourBinInstructional.C:194 FourBinInstructional.C:195 FourBinInstructional.C:196 FourBinInstructional.C:197 FourBinInstructional.C:198 FourBinInstructional.C:199 FourBinInstructional.C:200 FourBinInstructional.C:201 FourBinInstructional.C:202 FourBinInstructional.C:203 FourBinInstructional.C:204 FourBinInstructional.C:205 FourBinInstructional.C:206 FourBinInstructional.C:207 FourBinInstructional.C:208 FourBinInstructional.C:209 FourBinInstructional.C:210 FourBinInstructional.C:211 FourBinInstructional.C:212 FourBinInstructional.C:213 FourBinInstructional.C:214 FourBinInstructional.C:215 FourBinInstructional.C:216 FourBinInstructional.C:217 FourBinInstructional.C:218 FourBinInstructional.C:219 FourBinInstructional.C:220 FourBinInstructional.C:221 FourBinInstructional.C:222 FourBinInstructional.C:223 FourBinInstructional.C:224 FourBinInstructional.C:225 FourBinInstructional.C:226 FourBinInstructional.C:227 FourBinInstructional.C:228 FourBinInstructional.C:229 FourBinInstructional.C:230 FourBinInstructional.C:231 FourBinInstructional.C:232 FourBinInstructional.C:233 FourBinInstructional.C:234 FourBinInstructional.C:235 FourBinInstructional.C:236 FourBinInstructional.C:237 FourBinInstructional.C:238 FourBinInstructional.C:239 FourBinInstructional.C:240 FourBinInstructional.C:241 FourBinInstructional.C:242 FourBinInstructional.C:243 FourBinInstructional.C:244 FourBinInstructional.C:245 FourBinInstructional.C:246 FourBinInstructional.C:247 FourBinInstructional.C:248 FourBinInstructional.C:249 FourBinInstructional.C:250 FourBinInstructional.C:251 FourBinInstructional.C:252 FourBinInstructional.C:253 FourBinInstructional.C:254 FourBinInstructional.C:255 FourBinInstructional.C:256 FourBinInstructional.C:257 FourBinInstructional.C:258 FourBinInstructional.C:259 FourBinInstructional.C:260 FourBinInstructional.C:261 FourBinInstructional.C:262 FourBinInstructional.C:263 FourBinInstructional.C:264 FourBinInstructional.C:265 FourBinInstructional.C:266 FourBinInstructional.C:267 FourBinInstructional.C:268 FourBinInstructional.C:269 FourBinInstructional.C:270 FourBinInstructional.C:271 FourBinInstructional.C:272 FourBinInstructional.C:273 FourBinInstructional.C:274 FourBinInstructional.C:275 FourBinInstructional.C:276 FourBinInstructional.C:277 FourBinInstructional.C:278 FourBinInstructional.C:279 FourBinInstructional.C:280 FourBinInstructional.C:281 FourBinInstructional.C:282 FourBinInstructional.C:283 FourBinInstructional.C:284 FourBinInstructional.C:285 FourBinInstructional.C:286 FourBinInstructional.C:287 FourBinInstructional.C:288 FourBinInstructional.C:289 FourBinInstructional.C:290 FourBinInstructional.C:291 FourBinInstructional.C:292 FourBinInstructional.C:293 FourBinInstructional.C:294 FourBinInstructional.C:295 FourBinInstructional.C:296 FourBinInstructional.C:297 FourBinInstructional.C:298 FourBinInstructional.C:299 FourBinInstructional.C:300 FourBinInstructional.C:301 FourBinInstructional.C:302 FourBinInstructional.C:303 FourBinInstructional.C:304 FourBinInstructional.C:305 FourBinInstructional.C:306 FourBinInstructional.C:307 FourBinInstructional.C:308 FourBinInstructional.C:309 FourBinInstructional.C:310 FourBinInstructional.C:311 FourBinInstructional.C:312 FourBinInstructional.C:313 FourBinInstructional.C:314 FourBinInstructional.C:315 FourBinInstructional.C:316 FourBinInstructional.C:317 FourBinInstructional.C:318 FourBinInstructional.C:319 FourBinInstructional.C:320 FourBinInstructional.C:321 FourBinInstructional.C:322 FourBinInstructional.C:323 FourBinInstructional.C:324 FourBinInstructional.C:325 FourBinInstructional.C:326 FourBinInstructional.C:327 FourBinInstructional.C:328 FourBinInstructional.C:329 FourBinInstructional.C:330 FourBinInstructional.C:331 FourBinInstructional.C:332 |
|