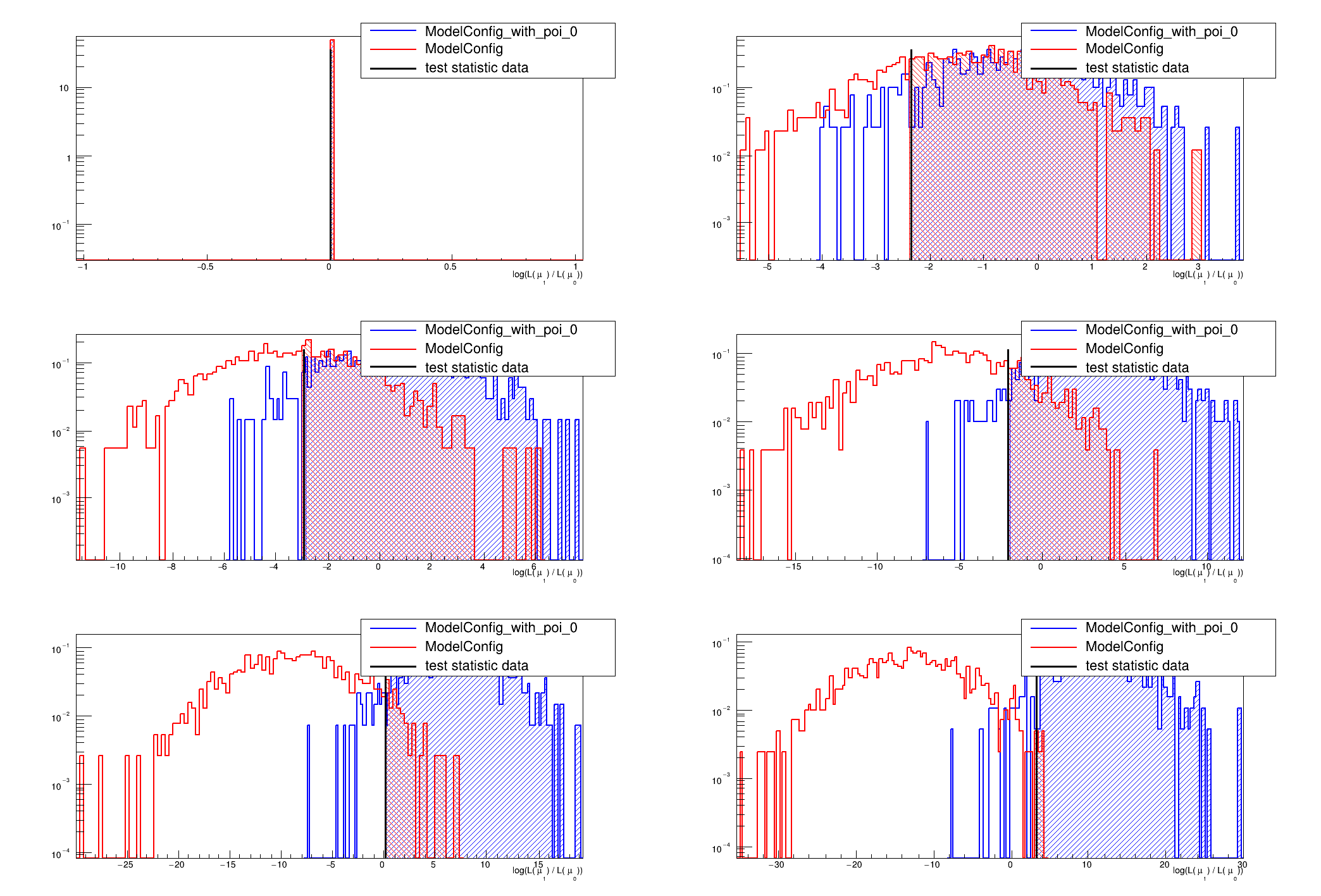
This macro will perform a scan of the p-values for computing the interval or limit
␛[1mRooFit v3.60 -- Developed by Wouter Verkerke and David Kirkby␛[0m
Copyright (C) 2000-2013 NIKHEF, University of California & Stanford University
All rights reserved, please read http://roofit.sourceforge.net/license.txt
0x5636e30dce40 results/example_combined_GaussExample_model.root
Running HypoTestInverter on the workspace combined
RooWorkspace(combined) combined contents
variables
---------
(Lumi,SigXsecOverSM,alpha_syst1,alpha_syst2,alpha_syst3,binWidth_obs_x_channel1_0,binWidth_obs_x_channel1_1,binWidth_obs_x_channel1_2,channelCat,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1,nom_alpha_syst1,nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1,nominalLumi,obs_x_channel1,weightVar)
p.d.f.s
-------
RooGaussian::alpha_syst1Constraint[ x=alpha_syst1 mean=nom_alpha_syst1 sigma=1 ] = 1
RooGaussian::alpha_syst2Constraint[ x=alpha_syst2 mean=nom_alpha_syst2 sigma=1 ] = 1
RooGaussian::alpha_syst3Constraint[ x=alpha_syst3 mean=nom_alpha_syst3 sigma=1 ] = 1
RooRealSumPdf::channel1_model[ binWidth_obs_x_channel1_0 * L_x_signal_channel1_overallSyst_x_Exp + binWidth_obs_x_channel1_1 * L_x_background1_channel1_overallSyst_x_StatUncert + binWidth_obs_x_channel1_2 * L_x_background2_channel1_overallSyst_x_StatUncert ] = 220
RooPoisson::gamma_stat_channel1_bin_0_constraint[ x=nom_gamma_stat_channel1_bin_0 mean=gamma_stat_channel1_bin_0_poisMean ] = 0.019943
RooPoisson::gamma_stat_channel1_bin_1_constraint[ x=nom_gamma_stat_channel1_bin_1 mean=gamma_stat_channel1_bin_1_poisMean ] = 0.039861
RooGaussian::lumiConstraint[ x=Lumi mean=nominalLumi sigma=0.1 ] = 1
RooProdPdf::model_channel1[ lumiConstraint * alpha_syst1Constraint * alpha_syst2Constraint * alpha_syst3Constraint * gamma_stat_channel1_bin_0_constraint * gamma_stat_channel1_bin_1_constraint * channel1_model(obs_x_channel1) ] = 0.174888
RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.174888
functions
--------
RooProduct::L_x_background1_channel1_overallSyst_x_StatUncert[ Lumi * background1_channel1_overallSyst_x_StatUncert ] = 0
RooProduct::L_x_background2_channel1_overallSyst_x_StatUncert[ Lumi * background2_channel1_overallSyst_x_StatUncert ] = 100
RooProduct::L_x_signal_channel1_overallSyst_x_Exp[ Lumi * signal_channel1_overallSyst_x_Exp ] = 10
RooStats::HistFactory::FlexibleInterpVar::background1_channel1_epsilon[ paramList=(alpha_syst2) ] = 1
RooHistFunc::background1_channel1_nominal[ depList=(obs_x_channel1) depList=(obs_x_channel1) ] = 0
RooProduct::background1_channel1_overallSyst_x_Exp[ background1_channel1_nominal * background1_channel1_epsilon ] = 0
RooProduct::background1_channel1_overallSyst_x_StatUncert[ mc_stat_channel1 * background1_channel1_overallSyst_x_Exp ] = 0
RooStats::HistFactory::FlexibleInterpVar::background2_channel1_epsilon[ paramList=(alpha_syst3) ] = 1
RooHistFunc::background2_channel1_nominal[ depList=(obs_x_channel1) depList=(obs_x_channel1) ] = 100
RooProduct::background2_channel1_overallSyst_x_Exp[ background2_channel1_nominal * background2_channel1_epsilon ] = 100
RooProduct::background2_channel1_overallSyst_x_StatUncert[ mc_stat_channel1 * background2_channel1_overallSyst_x_Exp ] = 100
RooProduct::gamma_stat_channel1_bin_0_poisMean[ gamma_stat_channel1_bin_0 * gamma_stat_channel1_bin_0_tau ] = 400
RooProduct::gamma_stat_channel1_bin_1_poisMean[ gamma_stat_channel1_bin_1 * gamma_stat_channel1_bin_1_tau ] = 100
ParamHistFunc::mc_stat_channel1[ ] = 1
RooStats::HistFactory::FlexibleInterpVar::signal_channel1_epsilon[ paramList=(alpha_syst1) ] = 1
RooHistFunc::signal_channel1_nominal[ depList=(obs_x_channel1) depList=(obs_x_channel1) ] = 10
RooProduct::signal_channel1_overallNorm_x_sigma_epsilon[ SigXsecOverSM * signal_channel1_epsilon ] = 1
RooProduct::signal_channel1_overallSyst_x_Exp[ signal_channel1_nominal * signal_channel1_overallNorm_x_sigma_epsilon ] = 10
datasets
--------
RooDataSet::asimovData(obs_x_channel1,weightVar,channelCat)
RooDataSet::obsData(channelCat,obs_x_channel1)
embedded datasets (in pdfs and functions)
-----------------------------------------
RooDataHist::signal_channel1nominalDHist(obs_x_channel1)
RooDataHist::background1_channel1nominalDHist(obs_x_channel1)
RooDataHist::background2_channel1nominalDHist(obs_x_channel1)
parameter snapshots
-------------------
NominalParamValues = (nom_alpha_syst2=0[C],nom_alpha_syst3=0[C],nom_gamma_stat_channel1_bin_0=400[C],nom_gamma_stat_channel1_bin_1=100[C],weightVar=0,obs_x_channel1=1.75,Lumi=1[C],nominalLumi=1[C],alpha_syst1=0[C],nom_alpha_syst1=0[C],alpha_syst2=0,alpha_syst3=0,gamma_stat_channel1_bin_0=1 +/- 0.05,gamma_stat_channel1_bin_1=1 +/- 0.1,SigXsecOverSM=1,binWidth_obs_x_channel1_0=2[C],binWidth_obs_x_channel1_1=2[C],binWidth_obs_x_channel1_2=2[C])
named sets
----------
ModelConfig_GlobalObservables:(nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
ModelConfig_NuisParams:(alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
ModelConfig_Observables:(obs_x_channel1,weightVar,channelCat)
ModelConfig_POI:(SigXsecOverSM)
globalObservables:(nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
observables:(obs_x_channel1,weightVar,channelCat)
generic objects
---------------
RooStats::ModelConfig::ModelConfig
Using data set obsData
StandardHypoTestInvDemo : POI initial value: SigXsecOverSM = 1
[#1] INFO:InputArguments -- HypoTestInverter ---- Input models:
using as S+B (null) model : ModelConfig
using as B (alternate) model : ModelConfig_with_poi_0
Doing a fixed scan in interval : 0 , 5
[#1] INFO:Eval -- HypoTestInverter::GetInterval - run a fixed scan
[#0] WARNING:InputArguments -- HypoTestInverter::RunFixedScan - xMax > upper bound, using xmax = 3
[#1] INFO:ObjectHandling -- RooWorkspace::saveSnaphot(combined) replacing previous snapshot with name ModelConfig__snapshot
[#0] PROGRESS:Eval -- Running for SigXsecOverSM = 0
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.158989
Snapshot:
1) 0x5636e3958370 RooRealVar:: SigXsecOverSM = 0 L(0 - 3) "SigXsecOverSM"
=== Using the following for ModelConfig_with_poi_0 ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.158989
Snapshot:
1) 0x5636e3909dc0 RooRealVar:: SigXsecOverSM = 0 L(0 - 3) "SigXsecOverSM"
[#0] PROGRESS:Generation -- Test Statistic on data: 0
[#1] INFO:InputArguments -- Profiling conditional MLEs for Null.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Null.
[#0] PROGRESS:Generation -- generated toys: 500 / 1000
[#1] INFO:InputArguments -- Profiling conditional MLEs for Alt.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Alt.
[#0] PROGRESS:Eval -- P values for SigXsecOverSM = 0
CLs = 1 +/- 0
CLb = 1 +/- 0
CLsplusb = 1 +/- 0
[#1] INFO:ObjectHandling -- RooWorkspace::saveSnaphot(combined) replacing previous snapshot with name ModelConfig__snapshot
[#0] PROGRESS:Eval -- Running for SigXsecOverSM = 0.6
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.168529
Snapshot:
1) 0x5636e3a0eff0 RooRealVar:: SigXsecOverSM = 0.6 L(0 - 3) "SigXsecOverSM"
=== Using the following for ModelConfig_with_poi_0 ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.168529
Snapshot:
1) 0x5636e395c990 RooRealVar:: SigXsecOverSM = 0 L(0 - 3) "SigXsecOverSM"
[#0] PROGRESS:Generation -- Test Statistic on data: -2.35222
[#1] INFO:InputArguments -- Profiling conditional MLEs for Null.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Null.
[#0] PROGRESS:Generation -- generated toys: 500 / 1000
[#1] INFO:InputArguments -- Profiling conditional MLEs for Alt.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Alt.
[#0] PROGRESS:Eval -- P values for SigXsecOverSM = 0.6
CLs = 0.819079 +/- 0.0188863
CLb = 0.912 +/- 0.0126693
CLsplusb = 0.747 +/- 0.0137474
[#1] INFO:ObjectHandling -- RooWorkspace::saveSnaphot(combined) replacing previous snapshot with name ModelConfig__snapshot
[#0] PROGRESS:Eval -- Running for SigXsecOverSM = 1.2
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.178068
Snapshot:
1) 0x5636e3aec300 RooRealVar:: SigXsecOverSM = 1.2 L(0 - 3) "SigXsecOverSM"
=== Using the following for ModelConfig_with_poi_0 ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.178068
Snapshot:
1) 0x5636e3a15050 RooRealVar:: SigXsecOverSM = 0 L(0 - 3) "SigXsecOverSM"
[#0] PROGRESS:Generation -- Test Statistic on data: -2.9364
[#1] INFO:InputArguments -- Profiling conditional MLEs for Null.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Null.
[#0] PROGRESS:Generation -- generated toys: 500 / 1000
[#1] INFO:InputArguments -- Profiling conditional MLEs for Alt.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Alt.
[#0] PROGRESS:Eval -- P values for SigXsecOverSM = 1.2
CLs = 0.48617 +/- 0.0176356
CLb = 0.94 +/- 0.0106207
CLsplusb = 0.457 +/- 0.0157528
[#1] INFO:ObjectHandling -- RooWorkspace::saveSnaphot(combined) replacing previous snapshot with name ModelConfig__snapshot
[#0] PROGRESS:Eval -- Running for SigXsecOverSM = 1.8
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.187607
Snapshot:
1) 0x5636e3ae12a0 RooRealVar:: SigXsecOverSM = 1.8 L(0 - 3) "SigXsecOverSM"
=== Using the following for ModelConfig_with_poi_0 ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.187607
Snapshot:
1) 0x5636e3b047b0 RooRealVar:: SigXsecOverSM = 0 L(0 - 3) "SigXsecOverSM"
[#0] PROGRESS:Generation -- Test Statistic on data: -2.05075
[#1] INFO:InputArguments -- Profiling conditional MLEs for Null.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Null.
[#0] PROGRESS:Generation -- generated toys: 500 / 1000
[#1] INFO:InputArguments -- Profiling conditional MLEs for Alt.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Alt.
[#0] PROGRESS:Eval -- P values for SigXsecOverSM = 1.8
CLs = 0.190678 +/- 0.0130363
CLb = 0.944 +/- 0.0102824
CLsplusb = 0.18 +/- 0.0121491
[#1] INFO:ObjectHandling -- RooWorkspace::saveSnaphot(combined) replacing previous snapshot with name ModelConfig__snapshot
[#0] PROGRESS:Eval -- Running for SigXsecOverSM = 2.4
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.197147
Snapshot:
1) 0x5636e3ad2bb0 RooRealVar:: SigXsecOverSM = 2.4 L(0 - 3) "SigXsecOverSM"
=== Using the following for ModelConfig_with_poi_0 ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.197147
Snapshot:
1) 0x5636e52500d0 RooRealVar:: SigXsecOverSM = 0 L(0 - 3) "SigXsecOverSM"
[#0] PROGRESS:Generation -- Test Statistic on data: 0.0783908
[#1] INFO:InputArguments -- Profiling conditional MLEs for Null.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Null.
[#0] PROGRESS:Generation -- generated toys: 500 / 1000
[#1] INFO:InputArguments -- Profiling conditional MLEs for Alt.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Alt.
[#0] PROGRESS:Eval -- P values for SigXsecOverSM = 2.4
CLs = 0.0550847 +/- 0.00746178
CLb = 0.944 +/- 0.0102824
CLsplusb = 0.052 +/- 0.00702111
[#1] INFO:ObjectHandling -- RooWorkspace::saveSnaphot(combined) replacing previous snapshot with name ModelConfig__snapshot
[#0] PROGRESS:Eval -- Running for SigXsecOverSM = 3
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.206686
Snapshot:
1) 0x5636e3b41680 RooRealVar:: SigXsecOverSM = 3 L(0 - 3) "SigXsecOverSM"
=== Using the following for ModelConfig_with_poi_0 ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.206686
Snapshot:
1) 0x5636e3adb460 RooRealVar:: SigXsecOverSM = 0 L(0 - 3) "SigXsecOverSM"
[#0] PROGRESS:Generation -- Test Statistic on data: 3.27476
[#1] INFO:InputArguments -- Profiling conditional MLEs for Null.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Null.
[#0] PROGRESS:Generation -- generated toys: 500 / 1000
[#1] INFO:InputArguments -- Profiling conditional MLEs for Alt.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Alt.
[#0] PROGRESS:Eval -- P values for SigXsecOverSM = 3
CLs = 0.00535332 +/- 0.00238893
CLb = 0.934 +/- 0.0111035
CLsplusb = 0.005 +/- 0.00223047
Time to perform limit scan
Real time 0:00:14, CP time 14.040
The computed upper limit is: 2.46135 +/- 0.0596845
Expected upper limits, using the B (alternate) model :
expected limit (median) 1.60988
expected limit (-1 sig) 1.34011
expected limit (+1 sig) 2.09019
expected limit (-2 sig) 1.14968
expected limit (+2 sig) 2.79445
[#0] WARNING:Plotting -- Could not determine xmin and xmax of sampling distribution that was given to plot.
[#0] WARNING:Plotting -- Could not determine xmin and xmax of sampling distribution that was given to plot.
#include <cassert>
using namespace std;
struct HypoTestInvOptions {
bool plotHypoTestResult = true;
bool writeResult = true;
bool optimize = true;
bool useVectorStore = true;
bool generateBinned = false;
bool noSystematics = false;
double nToysRatio = 2;
double maxPOI = -1;
bool useProof = false;
int nworkers = 0;
bool enableDetailedOutput =
false;
bool rebuild = false;
int nToyToRebuild = 100;
int rebuildParamValues = 0;
int initialFit = -1;
int randomSeed = -1;
int nAsimovBins = 0;
bool reuseAltToys = false;
double confLevel = 0.95;
std::string minimizerType =
"";
std::string massValue = "";
int printLevel = 0;
bool useNLLOffset = false;
};
HypoTestInvOptions optHTInv;
class HypoTestInvTool {
public:
HypoTestInvTool();
~HypoTestInvTool(){};
HypoTestInverterResult *RunInverter(
RooWorkspace *w,
const char *modelSBName,
const char *modelBName,
const char *dataName,
int type,
int testStatType,
bool useCLs,
int npoints,
double poimin, double poimax, int ntoys, bool useNumberCounting = false,
const char *nuisPriorName = 0);
void AnalyzeResult(HypoTestInverterResult *
r,
int calculatorType,
int testStatType,
bool useCLs,
int npoints,
const char *fileNameBase = 0);
void SetParameter(
const char *
name,
const char *value);
void SetParameter(
const char *
name,
bool value);
void SetParameter(
const char *
name,
int value);
void SetParameter(
const char *
name,
double value);
private:
bool mPlotHypoTestResult;
bool mWriteResult;
bool mOptimize;
bool mUseVectorStore;
bool mGenerateBinned;
bool mUseProof;
bool mRebuild;
bool mReuseAltToys;
bool mEnableDetOutput;
int mNWorkers;
int mNToyToRebuild;
int mRebuildParamValues;
int mPrintLevel;
int mInitialFit;
int mRandomSeed;
double mNToysRatio;
double mMaxPoi;
int mAsimovBins;
std::string mMassValue;
std::string
mMinimizerType;
};
}
RooStats::HypoTestInvTool::HypoTestInvTool()
: mPlotHypoTestResult(true), mWriteResult(false), mOptimize(true), mUseVectorStore(true), mGenerateBinned(false),
mUseProof(false), mEnableDetOutput(false), mRebuild(false), mReuseAltToys(false), mNWorkers(4),
mNToyToRebuild(100), mRebuildParamValues(0), mPrintLevel(0), mInitialFit(-1), mRandomSeed(-1), mNToysRatio(2),
mMaxPoi(-1), mAsimovBins(0), mMassValue(""), mMinimizerType(""), mResultFileName()
{
}
void RooStats::HypoTestInvTool::SetParameter(
const char *
name,
bool value)
{
std::string s_name(
name);
if (s_name.find("PlotHypoTestResult") != std::string::npos)
mPlotHypoTestResult = value;
if (s_name.find("WriteResult") != std::string::npos)
mWriteResult = value;
if (s_name.find("Optimize") != std::string::npos)
mOptimize = value;
if (s_name.find("UseVectorStore") != std::string::npos)
mUseVectorStore = value;
if (s_name.find("GenerateBinned") != std::string::npos)
mGenerateBinned = value;
if (s_name.find("UseProof") != std::string::npos)
mUseProof = value;
if (s_name.find("EnableDetailedOutput") != std::string::npos)
mEnableDetOutput = value;
if (s_name.find("Rebuild") != std::string::npos)
mRebuild = value;
if (s_name.find("ReuseAltToys") != std::string::npos)
mReuseAltToys = value;
return;
}
void RooStats::HypoTestInvTool::SetParameter(
const char *
name,
int value)
{
std::string s_name(
name);
if (s_name.find("NWorkers") != std::string::npos)
mNWorkers = value;
if (s_name.find("NToyToRebuild") != std::string::npos)
mNToyToRebuild = value;
if (s_name.find("RebuildParamValues") != std::string::npos)
mRebuildParamValues = value;
if (s_name.find("PrintLevel") != std::string::npos)
mPrintLevel = value;
if (s_name.find("InitialFit") != std::string::npos)
mInitialFit = value;
if (s_name.find("RandomSeed") != std::string::npos)
mRandomSeed = value;
if (s_name.find("AsimovBins") != std::string::npos)
mAsimovBins = value;
return;
}
void RooStats::HypoTestInvTool::SetParameter(
const char *
name,
double value)
{
std::string s_name(
name);
if (s_name.find("NToysRatio") != std::string::npos)
mNToysRatio = value;
if (s_name.find("MaxPOI") != std::string::npos)
mMaxPoi = value;
return;
}
void RooStats::HypoTestInvTool::SetParameter(
const char *
name,
const char *value)
{
std::string s_name(
name);
if (s_name.find("MassValue") != std::string::npos)
mMassValue.assign(value);
if (s_name.find("MinimizerType") != std::string::npos)
mMinimizerType.assign(value);
if (s_name.find("ResultFileName") != std::string::npos)
mResultFileName = value;
return;
}
void StandardHypoTestInvDemo(const char *infile = 0, const char *wsName = "combined",
const char *modelSBName = "ModelConfig", const char *modelBName = "",
const char *dataName = "obsData", int calculatorType = 0, int testStatType = 0,
bool useCLs = true, int npoints = 6, double poimin = 0, double poimax = 5,
int ntoys = 1000, bool useNumberCounting = false, const char *nuisPriorName = 0)
{
if (filename.IsNull()) {
filename = "results/example_combined_GaussExample_model.root";
if (!fileExist) {
#ifdef _WIN32
cout << "HistFactory file cannot be generated on Windows - exit" << endl;
return;
#endif
cout << "will run standard hist2workspace example" << endl;
gROOT->ProcessLine(
".! prepareHistFactory .");
gROOT->ProcessLine(
".! hist2workspace config/example.xml");
cout << "\n\n---------------------" << endl;
cout << "Done creating example input" << endl;
cout << "---------------------\n\n" << endl;
}
} else
filename = infile;
cout << "StandardRooStatsDemoMacro: Input file " << filename << " is not found" << endl;
return;
}
HypoTestInvTool calc;
calc.SetParameter("PlotHypoTestResult", optHTInv.plotHypoTestResult);
calc.SetParameter("WriteResult", optHTInv.writeResult);
calc.SetParameter("Optimize", optHTInv.optimize);
calc.SetParameter("UseVectorStore", optHTInv.useVectorStore);
calc.SetParameter("GenerateBinned", optHTInv.generateBinned);
calc.SetParameter("NToysRatio", optHTInv.nToysRatio);
calc.SetParameter("MaxPOI", optHTInv.maxPOI);
calc.SetParameter("UseProof", optHTInv.useProof);
calc.SetParameter("EnableDetailedOutput", optHTInv.enableDetailedOutput);
calc.SetParameter("NWorkers", optHTInv.nworkers);
calc.SetParameter("Rebuild", optHTInv.rebuild);
calc.SetParameter("ReuseAltToys", optHTInv.reuseAltToys);
calc.SetParameter("NToyToRebuild", optHTInv.nToyToRebuild);
calc.SetParameter("RebuildParamValues", optHTInv.rebuildParamValues);
calc.SetParameter("MassValue", optHTInv.massValue.c_str());
calc.SetParameter("MinimizerType", optHTInv.minimizerType.c_str());
calc.SetParameter("PrintLevel", optHTInv.printLevel);
calc.SetParameter("InitialFit", optHTInv.initialFit);
calc.SetParameter("ResultFileName", optHTInv.resultFileName);
calc.SetParameter("RandomSeed", optHTInv.randomSeed);
calc.SetParameter("AsimovBins", optHTInv.nAsimovBins);
if (optHTInv.useNLLOffset)
HypoTestInverterResult *
r = 0;
std::cout << w << "\t" << filename << std::endl;
if (w != NULL) {
r = calc.RunInverter(w, modelSBName, modelBName, dataName, calculatorType, testStatType, useCLs, npoints, poimin,
poimax, ntoys, useNumberCounting, nuisPriorName);
std::cerr << "Error running the HypoTestInverter - Exit " << std::endl;
return;
}
} else {
std::cout << "Reading an HypoTestInverterResult with name " << wsName << " from file " << filename << std::endl;
r =
dynamic_cast<HypoTestInverterResult *
>(
file->Get(wsName));
std::cerr << "File " << filename << " does not contain a workspace or an HypoTestInverterResult - Exit "
<< std::endl;
return;
}
}
calc.AnalyzeResult(
r, calculatorType, testStatType, useCLs, npoints, infile);
return;
}
void RooStats::HypoTestInvTool::AnalyzeResult(HypoTestInverterResult *
r,
int calculatorType,
int testStatType,
bool useCLs, int npoints, const char *fileNameBase)
{
double lowerLimit = 0;
double llError = 0;
#if defined ROOT_SVN_VERSION && ROOT_SVN_VERSION >= 44126
lowerLimit =
r->LowerLimit();
llError =
r->LowerLimitEstimatedError();
}
#else
lowerLimit =
r->LowerLimit();
llError =
r->LowerLimitEstimatedError();
#endif
double upperLimit =
r->UpperLimit();
double ulError =
r->UpperLimitEstimatedError();
if (lowerLimit < upperLimit * (1. - 1.E-4) && lowerLimit != 0)
std::cout << "The computed lower limit is: " << lowerLimit << " +/- " << llError << std::endl;
std::cout << "The computed upper limit is: " << upperLimit << " +/- " << ulError << std::endl;
std::cout << "Expected upper limits, using the B (alternate) model : " << std::endl;
std::cout <<
" expected limit (median) " <<
r->GetExpectedUpperLimit(0) << std::endl;
std::cout <<
" expected limit (-1 sig) " <<
r->GetExpectedUpperLimit(-1) << std::endl;
std::cout <<
" expected limit (+1 sig) " <<
r->GetExpectedUpperLimit(1) << std::endl;
std::cout <<
" expected limit (-2 sig) " <<
r->GetExpectedUpperLimit(-2) << std::endl;
std::cout <<
" expected limit (+2 sig) " <<
r->GetExpectedUpperLimit(2) << std::endl;
if (mEnableDetOutput) {
mWriteResult = true;
Info(
"StandardHypoTestInvDemo",
"detailed output will be written in output result file");
}
if (
r != NULL && mWriteResult) {
const char *calcType = (calculatorType == 0) ? "Freq" : (calculatorType == 1) ? "Hybr" : "Asym";
const char *limitType = (useCLs) ? "CLs" : "Cls+b";
const char *scanType = (npoints < 0) ? "auto" : "grid";
if (mResultFileName.IsNull()) {
mResultFileName =
TString::Format(
"%s_%s_%s_ts%d_", calcType, limitType, scanType, testStatType);
if (mMassValue.size() > 0) {
mResultFileName += mMassValue.c_str();
mResultFileName += "_";
}
name.Replace(0,
name.Last(
'/') + 1,
"");
}
TString uldistFile =
"RULDist.root";
if (existULDist) {
if (fileULDist)
ulDist = fileULDist->
Get(
"RULDist");
}
TFile *fileOut =
new TFile(mResultFileName,
"RECREATE");
if (ulDist)
Info(
"StandardHypoTestInvDemo",
"HypoTestInverterResult has been written in the file %s", mResultFileName.Data());
}
std::string typeName = "";
if (calculatorType == 0)
typeName = "Frequentist";
if (calculatorType == 1)
typeName = "Hybrid";
else if (calculatorType == 2 || calculatorType == 3) {
typeName = "Asymptotic";
mPlotHypoTestResult = false;
}
HypoTestInverterPlot *plot =
new HypoTestInverterPlot(
"HTI_Result_Plot", plotTitle,
r);
plot->Draw("CLb 2CL");
const int nEntries =
r->ArraySize();
if (mPlotHypoTestResult) {
if (nEntries > 1) {
}
for (int i = 0; i < nEntries; i++) {
if (nEntries > 1)
SamplingDistPlot *pl = plot->MakeTestStatPlot(i);
pl->SetLogYaxis(true);
pl->Draw();
}
}
}
HypoTestInverterResult *RooStats::HypoTestInvTool::RunInverter(
RooWorkspace *w,
const char *modelSBName,
const char *modelBName,
const char *dataName,
int type,
int testStatType, bool useCLs, int npoints,
double poimin, double poimax, int ntoys,
bool useNumberCounting, const char *nuisPriorName)
{
std::cout <<
"Running HypoTestInverter on the workspace " << w->
GetName() << std::endl;
if (!data) {
Error(
"StandardHypoTestDemo",
"Not existing data %s", dataName);
return 0;
} else
std::cout << "Using data set " << dataName << std::endl;
if (mUseVectorStore) {
}
ModelConfig *bModel = (ModelConfig *)w->
obj(modelBName);
ModelConfig *sbModel = (ModelConfig *)w->
obj(modelSBName);
if (!sbModel) {
Error(
"StandardHypoTestDemo",
"Not existing ModelConfig %s", modelSBName);
return 0;
}
if (!sbModel->GetPdf()) {
Error(
"StandardHypoTestDemo",
"Model %s has no pdf ", modelSBName);
return 0;
}
if (!sbModel->GetParametersOfInterest()) {
Error(
"StandardHypoTestDemo",
"Model %s has no poi ", modelSBName);
return 0;
}
if (!sbModel->GetObservables()) {
Error(
"StandardHypoTestInvDemo",
"Model %s has no observables ", modelSBName);
return 0;
}
if (!sbModel->GetSnapshot()) {
Info(
"StandardHypoTestInvDemo",
"Model %s has no snapshot - make one using model poi", modelSBName);
sbModel->SetSnapshot(*sbModel->GetParametersOfInterest());
}
if (optHTInv.noSystematics) {
const RooArgSet *nuisPar = sbModel->GetNuisanceParameters();
if (nuisPar && nuisPar->
getSize() > 0) {
std::cout << "StandardHypoTestInvDemo"
<< " - Switch off all systematics by setting them constant to their initial values" << std::endl;
}
if (bModel) {
const RooArgSet *bnuisPar = bModel->GetNuisanceParameters();
if (bnuisPar)
}
}
if (!bModel || bModel == sbModel) {
Info(
"StandardHypoTestInvDemo",
"The background model %s does not exist", modelBName);
Info(
"StandardHypoTestInvDemo",
"Copy it from ModelConfig %s and set POI to zero", modelSBName);
bModel = (ModelConfig *)sbModel->Clone();
if (!var)
return 0;
double oldval = var->
getVal();
} else {
if (!bModel->GetSnapshot()) {
Info(
"StandardHypoTestInvDemo",
"Model %s has no snapshot - make one using model poi and 0 values ",
modelBName);
if (var) {
double oldval = var->
getVal();
} else {
Error(
"StandardHypoTestInvDemo",
"Model %s has no valid poi", modelBName);
return 0;
}
}
}
bool hasNuisParam = (sbModel->GetNuisanceParameters() && sbModel->GetNuisanceParameters()->getSize() > 0);
bool hasGlobalObs = (sbModel->GetGlobalObservables() && sbModel->GetGlobalObservables()->getSize() > 0);
if (hasNuisParam && !hasGlobalObs) {
if (constrPdf) {
Warning(
"StandardHypoTestInvDemo",
"Model %s has nuisance parameters but no global observables associated",
sbModel->GetName());
"\tThe effect of the nuisance parameters will not be treated correctly ");
}
}
}
RooArgSet *allParams = sbModel->GetPdf()->getParameters(*data);
delete allParams;
const RooArgSet *poiSet = sbModel->GetParametersOfInterest();
std::cout <<
"StandardHypoTestInvDemo : POI initial value: " << poi->
GetName() <<
" = " << poi->
getVal()
<< std::endl;
bool doFit = mInitialFit;
if (testStatType == 0 && mInitialFit == -1)
doFit = false;
if (
type == 3 && mInitialFit == -1)
doFit = false;
double poihat = 0;
if (mMinimizerType.size() == 0)
else
Info(
"StandardHypoTestInvDemo",
"Using %s as minimizer for computing the test statistic",
if (doFit) {
Info(
"StandardHypoTestInvDemo",
" Doing a first fit to the observed data ");
if (sbModel->GetNuisanceParameters())
constrainParams.
add(*sbModel->GetNuisanceParameters());
"Fit to the model failed - try with strategy 1 and perform first an Hesse computation");
fitres = sbModel->GetPdf()->fitTo(
}
Warning(
"StandardHypoTestInvDemo",
" Fit still failed - continue anyway.....");
std::cout <<
"StandardHypoTestInvDemo - Best Fit value : " << poi->
GetName() <<
" = " << poihat <<
" +/- " std::cout << "Time for fitting : ";
sbModel->SetSnapshot(*sbModel->GetParametersOfInterest());
std::cout << "StandardHypoTestInvo: snapshot of S+B Model " << sbModel->GetName()
<< " is set to the best fit value" << std::endl;
}
if (testStatType == 0) {
if (!doFit)
Info(
"StandardHypoTestInvDemo",
"Using LEP test statistic - an initial fit is not done and the TS will use "
"the nuisances at the model value");
else
Info(
"StandardHypoTestInvDemo",
"Using LEP test statistic - an initial fit has been done and the TS will use "
"the nuisances at the best fit value");
}
SimpleLikelihoodRatioTestStat slrts(*sbModel->GetPdf(), *bModel->GetPdf());
RooArgSet nullParams(*sbModel->GetSnapshot());
if (sbModel->GetNuisanceParameters())
nullParams.add(*sbModel->GetNuisanceParameters());
if (sbModel->GetSnapshot())
slrts.SetNullParameters(nullParams);
if (bModel->GetNuisanceParameters())
altParams.add(*bModel->GetNuisanceParameters());
if (bModel->GetSnapshot())
slrts.SetAltParameters(altParams);
if (mEnableDetOutput)
slrts.EnableDetailedOutput();
RatioOfProfiledLikelihoodsTestStat ropl(*sbModel->GetPdf(), *bModel->GetPdf(), bModel->GetSnapshot());
ropl.SetSubtractMLE(false);
if (testStatType == 11)
ropl.SetSubtractMLE(true);
ropl.SetPrintLevel(mPrintLevel);
ropl.SetMinimizer(mMinimizerType.c_str());
if (mEnableDetOutput)
ropl.EnableDetailedOutput();
ProfileLikelihoodTestStat profll(*sbModel->GetPdf());
if (testStatType == 3)
profll.SetOneSided(true);
if (testStatType == 4)
profll.SetSigned(true);
profll.SetMinimizer(mMinimizerType.c_str());
profll.SetPrintLevel(mPrintLevel);
if (mEnableDetOutput)
profll.EnableDetailedOutput();
profll.SetReuseNLL(mOptimize);
slrts.SetReuseNLL(mOptimize);
ropl.SetReuseNLL(mOptimize);
if (mOptimize) {
profll.SetStrategy(0);
ropl.SetStrategy(0);
}
if (mMaxPoi > 0)
MaxLikelihoodEstimateTestStat maxll(*sbModel->GetPdf(), *poi);
NumEventsTestStat nevtts;
AsymptoticCalculator::SetPrintLevel(mPrintLevel);
HypoTestCalculatorGeneric *hc = 0;
hc = new FrequentistCalculator(*data, *bModel, *sbModel);
hc = new HybridCalculator(*data, *bModel, *sbModel);
hc = new AsymptoticCalculator(*data, *bModel, *sbModel, false);
hc = new AsymptoticCalculator(*data, *bModel, *sbModel,
true);
else {
Error(
"StandardHypoTestInvDemo",
"Invalid - calculator type = %d supported values are only :\n\t\t\t 0 "
"(Frequentist) , 1 (Hybrid) , 2 (Asymptotic) ",
return 0;
}
TestStatistic *testStat = 0;
if (testStatType == 0)
testStat = &slrts;
if (testStatType == 1 || testStatType == 11)
testStat = &ropl;
if (testStatType == 2 || testStatType == 3 || testStatType == 4)
testStat = &profll;
if (testStatType == 5)
testStat = &maxll;
if (testStatType == 6)
testStat = &nevtts;
if (testStat == 0) {
Error(
"StandardHypoTestInvDemo",
"Invalid - test statistic type = %d supported values are only :\n\t\t\t 0 (SLR) "
", 1 (Tevatron) , 2 (PLR), 3 (PLR1), 4(MLE)",
testStatType);
return 0;
}
ToyMCSampler *toymcs = (ToyMCSampler *)hc->GetTestStatSampler();
if (toymcs && (
type == 0 ||
type == 1)) {
if (sbModel->GetPdf()->canBeExtended()) {
if (useNumberCounting)
Warning(
"StandardHypoTestInvDemo",
"Pdf is extended: but number counting flag is set: ignore it ");
} else {
if (!useNumberCounting) {
Info(
"StandardHypoTestInvDemo",
"Pdf is not extended: number of events to generate taken from observed data set is %d", nEvents);
toymcs->SetNEventsPerToy(nEvents);
} else {
Info(
"StandardHypoTestInvDemo",
"using a number counting pdf");
toymcs->SetNEventsPerToy(1);
}
}
toymcs->SetTestStatistic(testStat);
Info(
"StandardHypoTestInvDemo",
"Data set is weighted, nentries = %d and sum of weights = %8.1f but toy "
"generation is unbinned - it would be faster to set mGenerateBinned to true\n",
}
toymcs->SetGenerateBinned(mGenerateBinned);
toymcs->SetUseMultiGen(mOptimize);
if (mGenerateBinned && sbModel->GetObservables()->getSize() > 2) {
Warning(
"StandardHypoTestInvDemo",
"generate binned is activated but the number of observable is %d. Too much "
"memory could be needed for allocating all the bins",
sbModel->GetObservables()->getSize());
}
if (mRandomSeed >= 0)
}
if (mReuseAltToys) {
hc->UseSameAltToys();
}
HybridCalculator *hhc = dynamic_cast<HybridCalculator *>(hc);
assert(hhc);
hhc->SetToys(ntoys, ntoys / mNToysRatio);
if (bModel->GetNuisanceParameters() || sbModel->GetNuisanceParameters()) {
toymcs->SetUseMultiGen(false);
ToyMCSampler::SetAlwaysUseMultiGen(false);
if (nuisPriorName)
nuisPdf = w->
pdf(nuisPriorName);
if (!nuisPdf) {
Info(
"StandardHypoTestInvDemo",
"No nuisance pdf given for the HybridCalculator - try to deduce pdf from the model");
if (bModel->GetPdf() && bModel->GetObservables())
else
}
if (!nuisPdf) {
if (bModel->GetPriorPdf()) {
nuisPdf = bModel->GetPriorPdf();
Info(
"StandardHypoTestInvDemo",
"No nuisance pdf given - try to use %s that is defined as a prior pdf in the B model",
} else {
Error(
"StandardHypoTestInvDemo",
"Cannot run Hybrid calculator because no prior on the nuisance "
"parameter is specified or can be derived");
return 0;
}
}
assert(nuisPdf);
Info(
"StandardHypoTestInvDemo",
"Using as nuisance Pdf ... ");
(bModel->GetNuisanceParameters()) ? bModel->GetNuisanceParameters() : sbModel->GetNuisanceParameters();
"Prior nuisance does not depend on nuisance parameters. They will be smeared in their full range");
}
delete np;
hhc->ForcePriorNuisanceAlt(*nuisPdf);
hhc->ForcePriorNuisanceNull(*nuisPdf);
}
if (testStatType == 3)
((AsymptoticCalculator *)hc)->SetOneSided(true);
if (testStatType != 2 && testStatType != 3)
"Only the PL test statistic can be used with AsymptoticCalculator - use by default a two-sided PL");
((FrequentistCalculator *)hc)->SetToys(ntoys, ntoys / mNToysRatio);
if (mEnableDetOutput)
((FrequentistCalculator *)hc)->StoreFitInfo(true);
((HybridCalculator *)hc)->SetToys(ntoys, ntoys / mNToysRatio);
}
HypoTestInverter calc(*hc);
calc.SetConfidenceLevel(optHTInv.confLevel);
calc.UseCLs(useCLs);
calc.SetVerbose(true);
if (mUseProof) {
ProofConfig
pc(*w, mNWorkers,
"",
kFALSE);
toymcs->SetProofConfig(&
pc);
}
if (npoints > 0) {
if (poimin > poimax) {
poimin = int(poihat);
poimax = int(poihat + 4 * poi->
getError());
}
std::cout << "Doing a fixed scan in interval : " << poimin << " , " << poimax << std::endl;
calc.SetFixedScan(npoints, poimin, poimax);
} else {
std::cout <<
"Doing an automatic scan in interval : " << poi->
getMin() <<
" , " << poi->
getMax() << std::endl;
}
HypoTestInverterResult *
r = calc.GetInterval();
std::cout << "Time to perform limit scan \n";
if (mRebuild) {
std::cout << "\n***************************************************************\n";
std::cout << "Rebuild the upper limit distribution by re-generating new set of pseudo-experiment and re-compute "
"for each of them a new upper limit\n\n";
allParams = sbModel->GetPdf()->getParameters(*data);
if (mRebuildParamValues != 0) {
*allParams = initialParameters;
}
if (mRebuildParamValues == 0 || mRebuildParamValues == 1) {
if (sbModel->GetNuisanceParameters())
constrainParams.
add(*sbModel->GetNuisanceParameters());
const RooArgSet *poiModel = sbModel->GetParametersOfInterest();
bModel->LoadSnapshot();
if (mRebuildParamValues == 0) {
std::cout << "rebuild using fitted parameter value for B-model snapshot" << std::endl;
constrainParams.
Print(
"v");
}
}
std::cout << "StandardHypoTestInvDemo: Initial parameters used for rebuilding: ";
delete allParams;
calc.SetCloseProof(1);
SamplingDistribution *limDist = calc.GetUpperLimitDistribution(true, mNToyToRebuild);
std::cout << "Time to rebuild distributions " << std::endl;
if (limDist) {
std::cout << "Expected limits after rebuild distribution " << std::endl;
std::cout << "expected upper limit (median of limit distribution) " << limDist->InverseCDF(0.5) << std::endl;
std::cout << "expected -1 sig limit (0.16% quantile of limit dist) "
std::cout << "expected +1 sig limit (0.84% quantile of limit dist) "
std::cout << "expected -2 sig limit (.025% quantile of limit dist) "
std::cout << "expected +2 sig limit (.975% quantile of limit dist) "
SamplingDistPlot limPlot((mNToyToRebuild < 200) ? 50 : 100);
limPlot.AddSamplingDistribution(limDist);
limPlot.GetTH1F()->SetStats(true);
limPlot.SetLineColor(
kBlue);
new TCanvas(
"limPlot",
"Upper Limit Distribution");
limPlot.Draw();
limDist->SetName("RULDist");
TFile *fileOut =
new TFile(
"RULDist.root",
"RECREATE");
limDist->Write();
} else
std::cout << "ERROR : failed to re-build distributions " << std::endl;
}
}
void ReadResult(const char *fileName, const char *resultName = "", bool useCLs = true)
{
StandardHypoTestInvDemo(fileName, resultName, "", "", "", 0, 0, useCLs);
}
#ifdef USE_AS_MAIN
{
StandardHypoTestInvDemo();
}
#endif
void Info(const char *location, const char *msgfmt,...)
void Error(const char *location, const char *msgfmt,...)
void Warning(const char *location, const char *msgfmt,...)
R__EXTERN TSystem * gSystem
static void SetDefaultMinimizer(const char *type, const char *algo=0)
static void SetDefaultStrategy(int strat)
static const std::string & DefaultMinimizerType()
RooArgSet * getObservables(const RooArgSet &set, Bool_t valueOnly=kTRUE) const
Return the observables of this pdf given a set of observables.
virtual void Print(Option_t *options=0) const
Print the object to the defaultPrintStream().
RooAbsArg * first() const
virtual void Print(Option_t *options=0) const
This method must be overridden when a class wants to print itself.
RooAbsData is the common abstract base class for binned and unbinned datasets.
static void setDefaultStorageType(StorageType s)
virtual Double_t sumEntries() const =0
virtual Bool_t isWeighted() const
void convertToVectorStore()
Convert tree-based storage to vector-based storage.
virtual Int_t numEntries() const
virtual Double_t getMax(const char *name=0) const
Get maximum of currently defined range.
virtual Double_t getMin(const char *name=0) const
Get miniminum of currently defined range.
Double_t getVal(const RooArgSet *normalisationSet=nullptr) const
Evaluate object.
RooArgSet is a container object that can hold multiple RooAbsArg objects.
RooArgSet * snapshot(bool deepCopy=true) const
Use RooAbsCollection::snapshot(), but return as RooArgSet.
virtual Bool_t add(const RooAbsCollection &col, Bool_t silent=kFALSE)
Add a collection of arguments to this collection by calling add() for each element in the source coll...
RooFitResult is a container class to hold the input and output of a PDF fit to a dataset.
static RooMsgService & instance()
Return reference to singleton instance.
StreamConfig & getStream(Int_t id)
static TRandom * randomGenerator()
Return a pointer to a singleton random-number generator implementation.
RooRealVar represents a variable that can be changed from the outside.
void setMax(const char *name, Double_t value)
Set maximum of name range to given value.
Double_t getError() const
virtual void setVal(Double_t value)
Set value of variable to 'value'.
The RooWorkspace is a persistable container for RooFit projects.
RooAbsData * data(const char *name) const
Retrieve dataset (binned or unbinned) with given name. A null pointer is returned if not found.
void Print(Option_t *opts=0) const
Print contents of the workspace.
TObject * obj(const char *name) const
Return any type of object (RooAbsArg, RooAbsData or generic object) with given name)
RooAbsPdf * pdf(const char *name) const
Retrieve p.d.f (RooAbsPdf) with given name. A null pointer is returned if not found.
TObject * Get(const char *namecycle) override
Return pointer to object identified by namecycle.
A ROOT file is a suite of consecutive data records (TKey instances) with a well defined format.
static TFile * Open(const char *name, Option_t *option="", const char *ftitle="", Int_t compress=ROOT::RCompressionSetting::EDefaults::kUseCompiledDefault, Int_t netopt=0)
Create / open a file.
void Close(Option_t *option="") override
Close a file.
virtual const char * GetName() const
Returns name of object.
Mother of all ROOT objects.
virtual Int_t Write(const char *name=0, Int_t option=0, Int_t bufsize=0)
Write this object to the current directory.
virtual const char * GetName() const
Returns name of object.
virtual void SetSeed(ULong_t seed=0)
Set the random generator seed.
void Start(Bool_t reset=kTRUE)
Start the stopwatch.
void Print(Option_t *option="") const
Print the real and cpu time passed between the start and stop events.
static TString Format(const char *fmt,...)
Static method which formats a string using a printf style format descriptor and return a TString.
virtual Bool_t AccessPathName(const char *path, EAccessMode mode=kFileExists)
Returns FALSE if one can access a file using the specified access mode.
double normal_cdf(double x, double sigma=1, double x0=0)
Cumulative distribution function of the normal (Gaussian) distribution (lower tail).
int main(int argc, char **argv)
The namespace RooFit contains mostly switches that change the behaviour of functions of PDFs (or othe...
RooCmdArg Constrain(const RooArgSet ¶ms)
RooCmdArg Strategy(Int_t code)
RooCmdArg Hesse(Bool_t flag=kTRUE)
RooCmdArg InitialHesse(Bool_t flag=kTRUE)
RooCmdArg Save(Bool_t flag=kTRUE)
RooCmdArg PrintLevel(Int_t code)
RooCmdArg Offset(Bool_t flag=kTRUE)
RooCmdArg Minimizer(const char *type, const char *alg=0)
Namespace for the RooStats classes.
bool SetAllConstant(const RooAbsCollection &coll, bool constant=true)
void RemoveConstantParameters(RooArgSet *set)
RooAbsPdf * MakeNuisancePdf(RooAbsPdf &pdf, const RooArgSet &observables, const char *name)
void UseNLLOffset(bool on)
Use an offset in NLL calculations.
bool IsNLLOffset()
Test of RooStats should by default offset NLL calculations.
void PrintListContent(const RooArgList &l, std::ostream &os=std::cout)
static constexpr double pc
Double_t Sqrt(Double_t x)
Int_t CeilNint(Double_t x)
void removeTopic(RooFit::MsgTopic oldTopic)

 Standard tutorial macro for performing an inverted hypothesis test for computing an interval
Standard tutorial macro for performing an inverted hypothesis test for computing an interval